db <- readr::read_csv(here("..","data","db_effect_sizes.csv")) %>%
mutate(across(c(C_n, Ex_n, C_mean, Ex_mean, C_SE, Ex_SE, C_SD, Ex_SD, lnRR, lnRR_var), as.numeric))%>%
mutate(across(where(is.character), as.factor))
db <- db %>%
mutate(Study = str_extract(Study_ID, ".*_\\d{4}"))Leave-one-out
To evaluate whether the overall meta-analytic estimate is driven by any single study, we perform a leave-one-out analysis at the study level.
For each iteration, we refit the full multilevel model after removing all effect sizes from one study and record the resulting overall lnRR estimate. If the pooled effect remains stable across iterations, this indicates that the main result is not unduly influenced by any single study.
VCV <- metafor::vcalc(
vi = lnRR_var,
cluster = Cohort_ID, # The clustering variable (Study_ID + Ex_ID)
obs = ES_ID, # The unique observation/effect size ID
rho = 0.5, # Assumed correlation between outcomes from the same cohort
data = db
)Because our model uses a variance–covariance matrix (VCV) to account for within-cohort dependence, we must recompute that matrix each time a study is removed. The vcalc_args object ensures that the same covariance structure (cluster variable and assumed correlation) is applied during each refitting step.
vcalc_args <- list(vi = "lnRR_var",
cluster = "Cohort_ID",
obs = "ES_ID",
rho = 0.5)ma_all <- rma.mv(yi = lnRR,
V = VCV,
random = list(~1 | Study_ID,
~1 | ES_ID,
~1 | Strain),
test = "t",
method = "REML",
data = db)
summary(ma_all)
Multivariate Meta-Analysis Model (k = 298; method: REML)
logLik Deviance AIC BIC AICc
-242.8851 485.7702 493.7702 508.5452 493.9072
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0494 0.2223 20 no Study_ID
sigma^2.2 0.2302 0.4798 298 no ES_ID
sigma^2.3 0.0000 0.0001 6 no Strain
Test for Heterogeneity:
Q(df = 297) = 5799.5331, p-val < .0001
Model Results:
estimate se tval df pval ci.lb ci.ub
-0.2120 0.0662 -3.2046 297 0.0015 -0.3422 -0.0818 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1loo_t<-leave_one_out(ma_all, group = "Study", vcalc_args =vcalc_args)loo_t name estimate lowerCL upperCL lowerPR upperPR
1 Krishnamurthy_2025 -0.1804714 -0.3067364 -0.05420638 -1.1310613 0.7701185
2 Freitas_2020 -0.2345608 -0.3662627 -0.10285889 -1.2898706 0.8207490
3 Saghari_2021 -0.2186383 -0.3569070 -0.08036970 -1.2871952 0.8499185
4 Escribano_2014 -0.1452226 -0.2177303 -0.07271486 -0.9724561 0.6820110
5 Papadakakis_2019 -0.2169011 -0.3567353 -0.07706679 -1.3045233 0.8707212
6 Fu_2023 -0.1993904 -0.3384132 -0.06036748 -1.2615261 0.8627454
7 Sampaio_2017 -0.2205727 -0.3622150 -0.07893040 -1.3101872 0.8690418
8 Fu_2025 -0.2184528 -0.3541891 -0.08271651 -1.2824768 0.8455712
9 Niehues_2011 -0.2185420 -0.3593437 -0.07774032 -1.2815548 0.8444707
10 Pangemanan_2024 -0.2078496 -0.3398487 -0.07585035 -1.2595242 0.8438251
11 Chen_2019 -0.2108857 -0.3478612 -0.07391022 -1.2811460 0.8593745
12 Ren_2024 -0.2076347 -0.3429355 -0.07233395 -1.2080585 0.7927891
13 Milbratz_2017 -0.2122834 -0.3504070 -0.07415990 -1.2868257 0.8622589
14 Flores_2018 -0.2227915 -0.3584293 -0.08715384 -1.2809836 0.8354005
15 Cheng_2024 -0.2126974 -0.3506180 -0.07477677 -1.2892410 0.8638462
16 Rizzolo_2021 -0.2132545 -0.3449213 -0.08158768 -1.2654004 0.8388914
17 Camargo_2013 -0.2126936 -0.3490734 -0.07631379 -1.2721668 0.8467797
18 Terzioglu_2020 -0.2175428 -0.3560578 -0.07902784 -1.2703522 0.8352666
19 Li_2010 -0.2216599 -0.3580481 -0.08527166 -1.2902300 0.8469103
20 Chikahisa_2007 -0.2220461 -0.3626064 -0.08148583 -1.3353737 0.8912814loo_plot<-orchard_leave1out(leave1out = loo_t,
xlab = "lnRR",
ylab = "Studies left out",
trunk.size = 0.3,
branch.size = 2,
alpha = 0.3,
legend.pos = "top.out")
loo_plot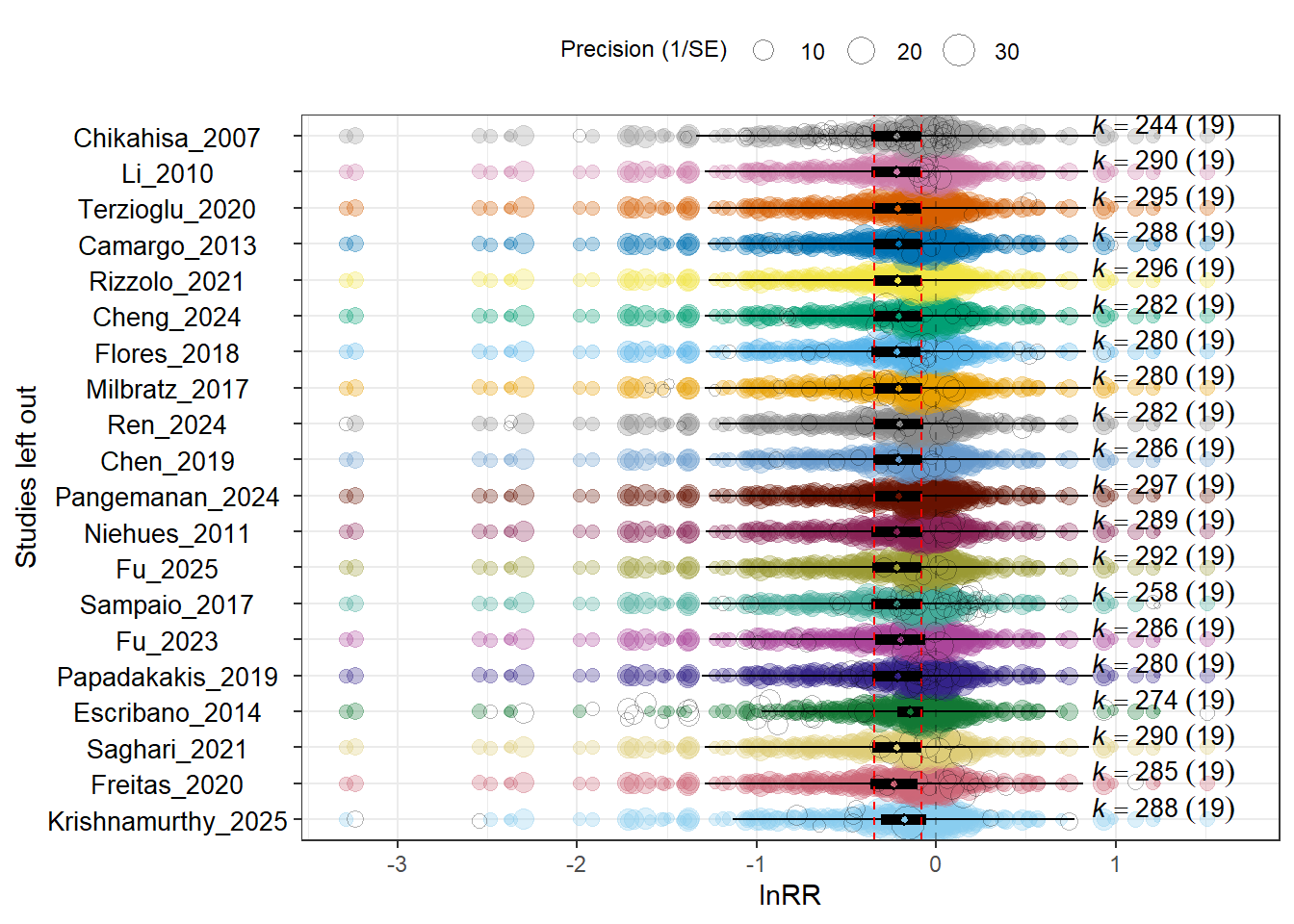
Across iterations, the pooled lnRR estimate remained within a narrow range and consistently below zero. No single study reversed the direction or substantially altered the magnitude of the overall effect. This indicates that the observed reduction in anxiety- and depression-like behaviors under music exposure is not driven by any individual dataset and is robust to study-level exclusion.
ggsave(filename = here("..","Plots", "loo_studies.pdf"),
plot = loo_plot,
dpi = 300, device = cairo_pdf,width = 12,
height = 10)
ggsave(filename = here("..","Plots", "loo_studies.jpg"),
plot = loo_plot,
width = 12,
height = 10,
dpi = 300)sessionInfo()R version 4.5.2 (2025-10-31 ucrt)
Platform: x86_64-w64-mingw32/x64
Running under: Windows 11 x64 (build 26200)
Matrix products: default
LAPACK version 3.12.1
locale:
[1] LC_COLLATE=English_Guernsey.utf8 LC_CTYPE=English_Guernsey.utf8
[3] LC_MONETARY=English_Guernsey.utf8 LC_NUMERIC=C
[5] LC_TIME=English_Guernsey.utf8
time zone: America/Edmonton
tzcode source: internal
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] orchaRd_2.1.3 metafor_4.8-0 numDeriv_2016.8-1.1
[4] metadat_1.4-0 Matrix_1.7-4 patchwork_1.3.2
[7] lubridate_1.9.4 forcats_1.0.1 stringr_1.6.0
[10] dplyr_1.1.4 purrr_1.2.1 readr_2.1.6
[13] tidyr_1.3.2 tibble_3.3.1 ggplot2_4.0.1
[16] tidyverse_2.0.0 knitr_1.51 here_1.0.2
[19] dtplyr_1.3.2 DT_0.34.0
loaded via a namespace (and not attached):
[1] beeswarm_0.4.0 gtable_0.3.6 xfun_0.56
[4] htmlwidgets_1.6.4 lattice_0.22-7 mathjaxr_2.0-0
[7] tzdb_0.5.0 vctrs_0.7.1 tools_4.5.2
[10] generics_0.1.4 parallel_4.5.2 sandwich_3.1-1
[13] pacman_0.5.1 pkgconfig_2.0.3 data.table_1.18.2.1
[16] RColorBrewer_1.1-3 S7_0.2.1 lifecycle_1.0.5
[19] compiler_4.5.2 farver_2.1.2 codetools_0.2-20
[22] vipor_0.4.7 htmltools_0.5.9 yaml_2.3.12
[25] pillar_1.11.1 crayon_1.5.3 MASS_7.3-65
[28] multcomp_1.4-29 nlme_3.1-168 tidyselect_1.2.1
[31] digest_0.6.39 mvtnorm_1.3-3 stringi_1.8.7
[34] labeling_0.4.3 splines_4.5.2 latex2exp_0.9.8
[37] rprojroot_2.1.1 fastmap_1.2.0 grid_4.5.2
[40] cli_3.6.5 magrittr_2.0.4 survival_3.8-6
[43] TH.data_1.1-5 withr_3.0.2 scales_1.4.0
[46] bit64_4.6.0-1 ggbeeswarm_0.7.3 timechange_0.3.0
[49] estimability_1.5.1 rmarkdown_2.30 emmeans_2.0.1
[52] bit_4.6.0 otel_0.2.0 zoo_1.8-15
[55] hms_1.1.4 coda_0.19-4.1 evaluate_1.0.5
[58] rlang_1.1.7 xtable_1.8-4 glue_1.8.0
[61] rstudioapi_0.18.0 vroom_1.7.0 jsonlite_2.0.0
[64] R6_2.6.1