pacman::p_load(
DT, dtplyr, here, knitr, tidyverse, patchwork, metafor, orchaRd, multcomp
)
db_all <- readr::read_csv(here("..","data","db_effect_sizes.csv")) %>%
mutate(across(where(is.character), as.factor)) %>%
mutate(across(c(lnRR, lnRR_var, lnVR, lnVR_var, lnCVR, lnCVR_var), as.numeric))%>%
mutate(
Control_conditions = as.character(Control_conditions),
Control_conditions = dplyr::recode(
Control_conditions,
"ambiet sound" = "Ambient sound",
"white noise" = "White noise"
),
Control_conditions = factor(Control_conditions)
)
# Per-metric datasets
db_rr <- db_all %>% filter(!is.na(lnRR), !is.na(lnRR_var))
db_vr <- db_all %>% filter(!is.na(lnVR), !is.na(lnVR_var))
db_cvr <- db_all %>% filter(!is.na(lnCVR), !is.na(lnCVR_var))
# Per-metric VCV matrices (rho = 0.5)
VCV_rr <- metafor::vcalc(vi = lnRR_var, cluster = Cohort_ID, obs = ES_ID, rho = 0.5, data = db_rr)
VCV_vr <- metafor::vcalc(vi = lnVR_var, cluster = Cohort_ID, obs = ES_ID, rho = 0.5, data = db_vr)
VCV_cvr<- metafor::vcalc(vi = lnCVR_var, cluster = Cohort_ID, obs = ES_ID, rho = 0.5, data = db_cvr)Uni-moderators
We fit uni-moderator multilevel meta-regressions to test whether study characteristics predict differences in:
the mean response (lnRR),
absolute variability (lnVR),
relative variability (lnCVR).
Each moderator is analysed separately (uni-moderator), using the same random-effects structure (Study_ID, ES_ID, Strain`) and a cohort-level VCV matrix (rho = 0.5). For categorical moderators, we drop levels with fewer than five effect sizes (k < 5) within each effect size metric, and subset the corresponding VCV matrix accordingly.
NoteHelper functions
Filter to common moderator levels (k ≥ 5 in each metric)
Code
filter_low_k_single <- function(dat, V, moderator, min_k = 5) {
mod_chr <- moderator
# If moderator is not a factor, just return as-is
if (!is.factor(dat[[mod_chr]])) {
return(list(data = dat, V = V, keep_levels = NULL, k = NULL))
}
k_counts <- dat %>%
count(.data[[mod_chr]], name = "k_es")
keep_levels <- k_counts %>%
filter(k_es >= min_k) %>%
pull(.data[[mod_chr]])
dat_f <- dat %>% filter(.data[[mod_chr]] %in% keep_levels)
idx <- which(dat[[mod_chr]] %in% keep_levels)
V_f <- V[idx, idx, drop = FALSE]
list(data = dat_f, V = V_f, keep_levels = keep_levels, k = k_counts)
}Fit one uni-moderator model + plot
Code
fit_unimod <- function(dat, V, yi, mods_formula, mod_name, xlab_expr) {
# Guard: if nothing left after filtering
if (nrow(dat) < 2) return(NULL)
mod <- metafor::rma.mv(
yi = dat[[yi]],
V = V,
mods = as.formula(mods_formula),
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain),
test = "t",
method = "REML",
sparse = TRUE,
data = dat
)
list(
data = dat,
model = mod,
r2 = round(orchaRd::r2_ml(mod), 4),
i2 = orchaRd::i2_ml(mod),
plot = orchard_plot(
mod,
mod = mod_name,
group = "Study_ID",
xlab = xlab_expr,
flip = TRUE,
legend.pos = "top.left",
trunk.size = 0.3,
branch.size = 2,
alpha = 0.3
) +
scale_colour_brewer(palette = "Set1") +
scale_fill_brewer(palette = "Dark2") +
theme(legend.direction = "vertical")
)
}Optional Tukey contrasts
Code
maybe_tukey <- function(dat, V, yi, moderator, min_k = 3) {
dat <- dat %>%
mutate("{moderator}" := droplevels(as.factor(.data[[moderator]])))
k <- nlevels(dat[[moderator]])
if (is.na(k) || k < min_k) return(NULL)
# Fit without intercept so coef = one per level
mods_formula <- as.formula(paste0("~ ", moderator, " - 1"))
m <- metafor::rma.mv(
yi = dat[[yi]],
V = V,
mods = mods_formula,
random = list(~1 | Study_ID,
~1 | ES_ID,
~1 | Strain),
test = "t",
method = "REML",
sparse = TRUE,
data = dat
)
cn <- names(coef(m))
K <- multcomp::contrMat(rep(1, k), type = "Tukey")
colnames(K) <- cn
g <- multcomp::glht(m, linfct = K)
base::summary(g, test = multcomp::adjusted("none"))
}Outcome type
MOD <- "Outcome_type"
MODS <- "~ Outcome_type"
f_rr <- filter_low_k_single(db_rr, VCV_rr, MOD, min_k = 5)
f_vr <- filter_low_k_single(db_vr, VCV_vr, MOD, min_k = 5)
f_cvr <- filter_low_k_single(db_cvr, VCV_cvr, MOD, min_k = 5)
res_rr <- fit_unimod(f_rr$data, f_rr$V, "lnRR", MODS, MOD, expression(lnRR))
res_vr <- fit_unimod(f_vr$data, f_vr$V, "lnVR", MODS, MOD, expression(lnVR))
res_cvr <- fit_unimod(f_cvr$data, f_cvr$V, "lnCVR", MODS, MOD, expression(lnCVR))lnRR
summary(res_rr$model)
Multivariate Meta-Analysis Model (k = 295; method: REML)
logLik Deviance AIC BIC AICc
-241.0967 482.1933 492.1933 510.5942 492.4024
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0495 0.2225 20 no Study_ID
sigma^2.2 0.2333 0.4830 295 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 293) = 5690.1059, p-val < .0001
Test of Moderators (coefficient 2):
F(df1 = 1, df2 = 293) = 0.0236, p-val = 0.8781
Model Results:
estimate se tval df pval ci.lb
intrcpt -0.2158 0.0710 -3.0389 293 0.0026 -0.3555
Outcome_typeDepression 0.0139 0.0907 0.1535 293 0.8781 -0.1645
ci.ub
intrcpt -0.0760 **
Outcome_typeDepression 0.1923
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_rr$r2 R2_marginal R2_conditional
0.0001 0.1752 res_rr$plot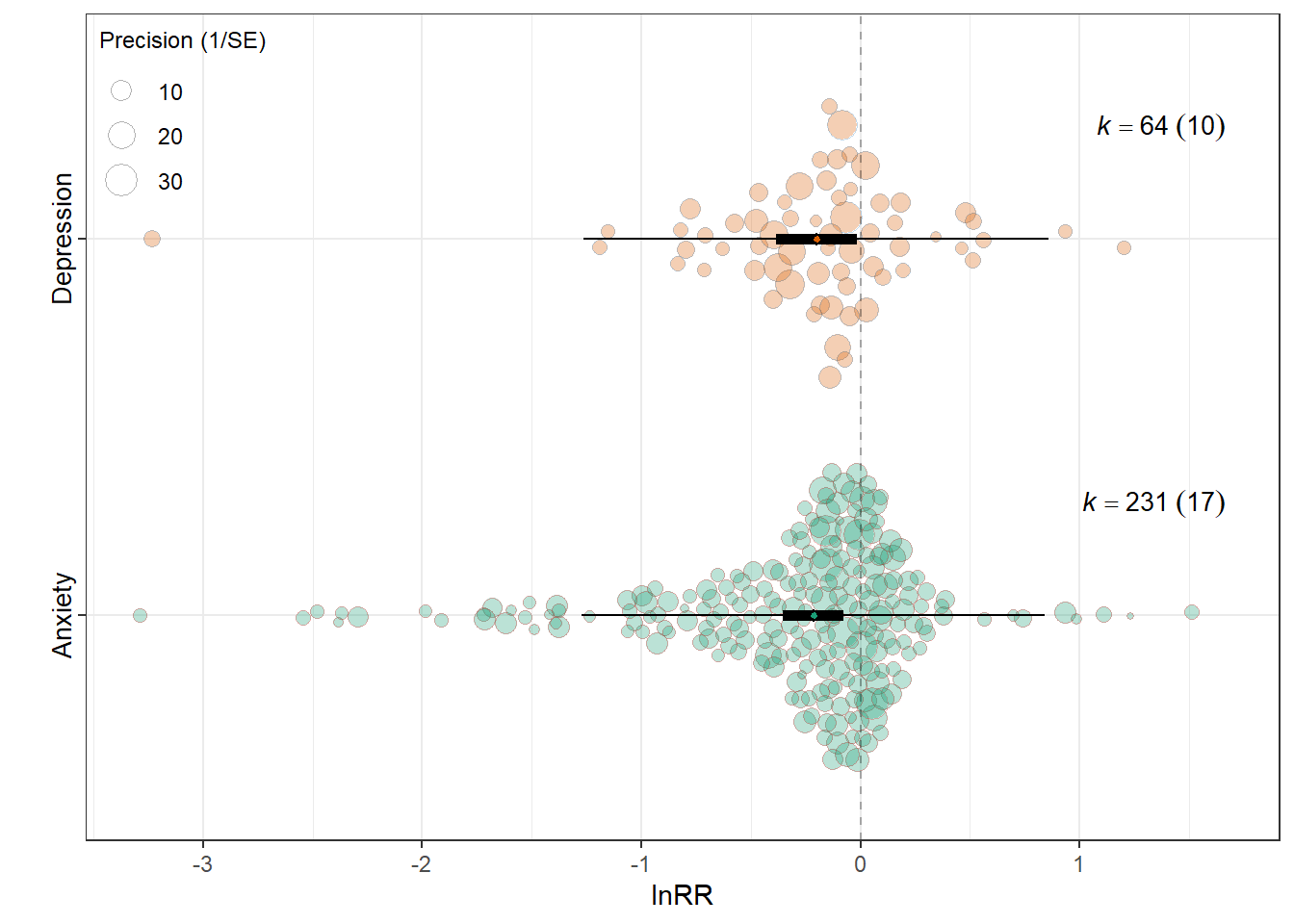
lnVR
summary(res_vr$model)
Multivariate Meta-Analysis Model (k = 219; method: REML)
logLik Deviance AIC BIC AICc
-339.3184 678.6368 688.6368 705.5363 688.9211
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.1310 0.3620 16 no Study_ID
sigma^2.2 1.1825 1.0874 219 no ES_ID
sigma^2.3 0.0000 0.0001 6 no Strain
Test for Residual Heterogeneity:
QE(df = 217) = 4218.8132, p-val < .0001
Test of Moderators (coefficient 2):
F(df1 = 1, df2 = 217) = 8.3073, p-val = 0.0043
Model Results:
estimate se tval df pval ci.lb
intrcpt 0.3112 0.1520 2.0473 217 0.0418 0.0116
Outcome_typeDepression -0.6401 0.2221 -2.8822 217 0.0043 -1.0779
ci.ub
intrcpt 0.6107 *
Outcome_typeDepression -0.2024 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_vr$r2 R2_marginal R2_conditional
0.0537 0.1481 res_vr$plot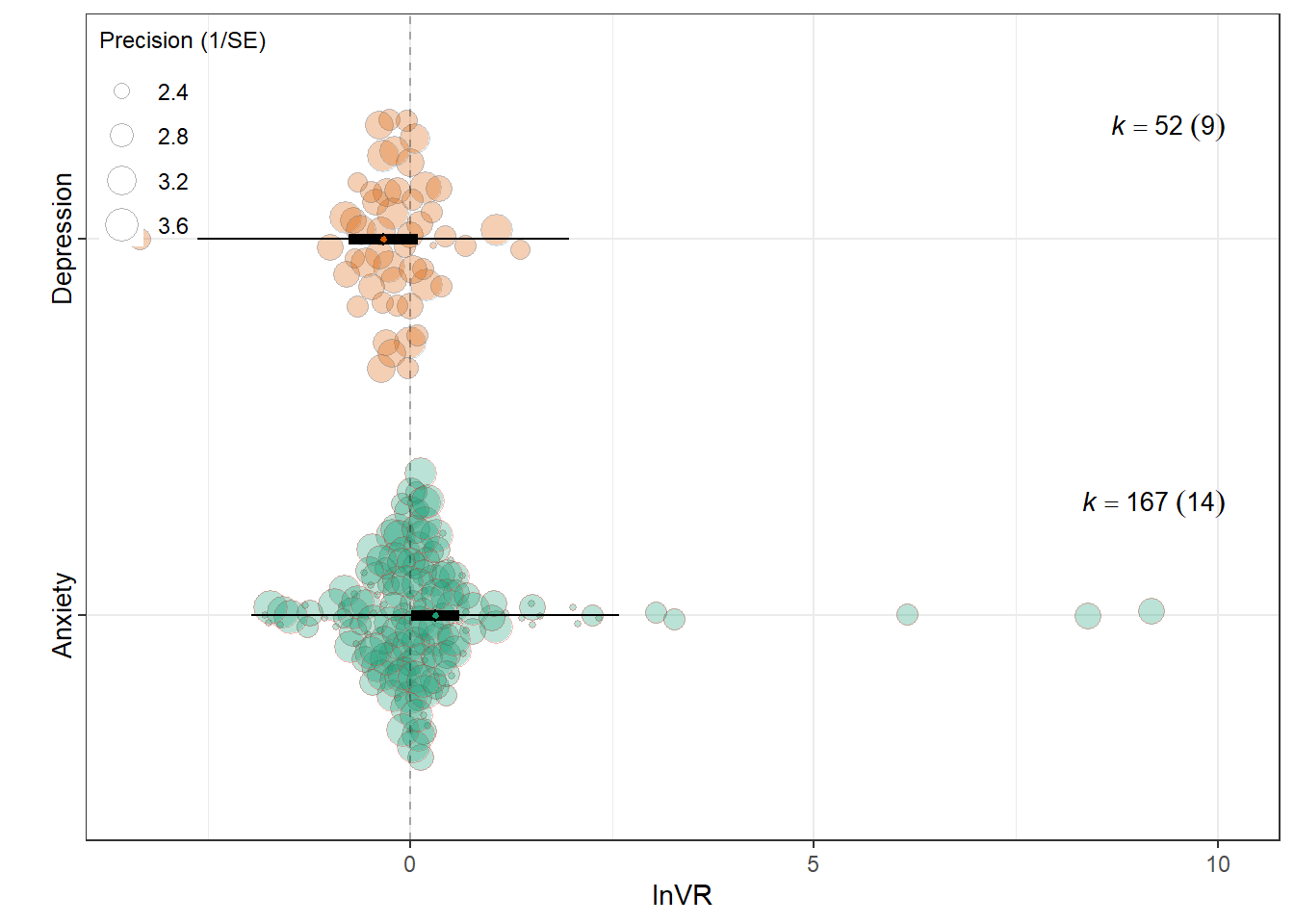
lnCVR
summary(res_cvr$model)
Multivariate Meta-Analysis Model (k = 219; method: REML)
logLik Deviance AIC BIC AICc
-279.5331 559.0662 569.0662 585.9656 569.3505
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.1050 0.3240 16 no Study_ID
sigma^2.2 0.5956 0.7718 219 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 217) = 1576.9857, p-val < .0001
Test of Moderators (coefficient 2):
F(df1 = 1, df2 = 217) = 1.1590, p-val = 0.2829
Model Results:
estimate se tval df pval ci.lb ci.ub
intrcpt 0.0615 0.1339 0.4590 217 0.6467 -0.2025 0.3255
Outcome_typeDepression -0.1830 0.1700 -1.0766 217 0.2829 -0.5180 0.1520
intrcpt
Outcome_typeDepression
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_cvr$r2 R2_marginal R2_conditional
0.0086 0.1572 res_cvr$plot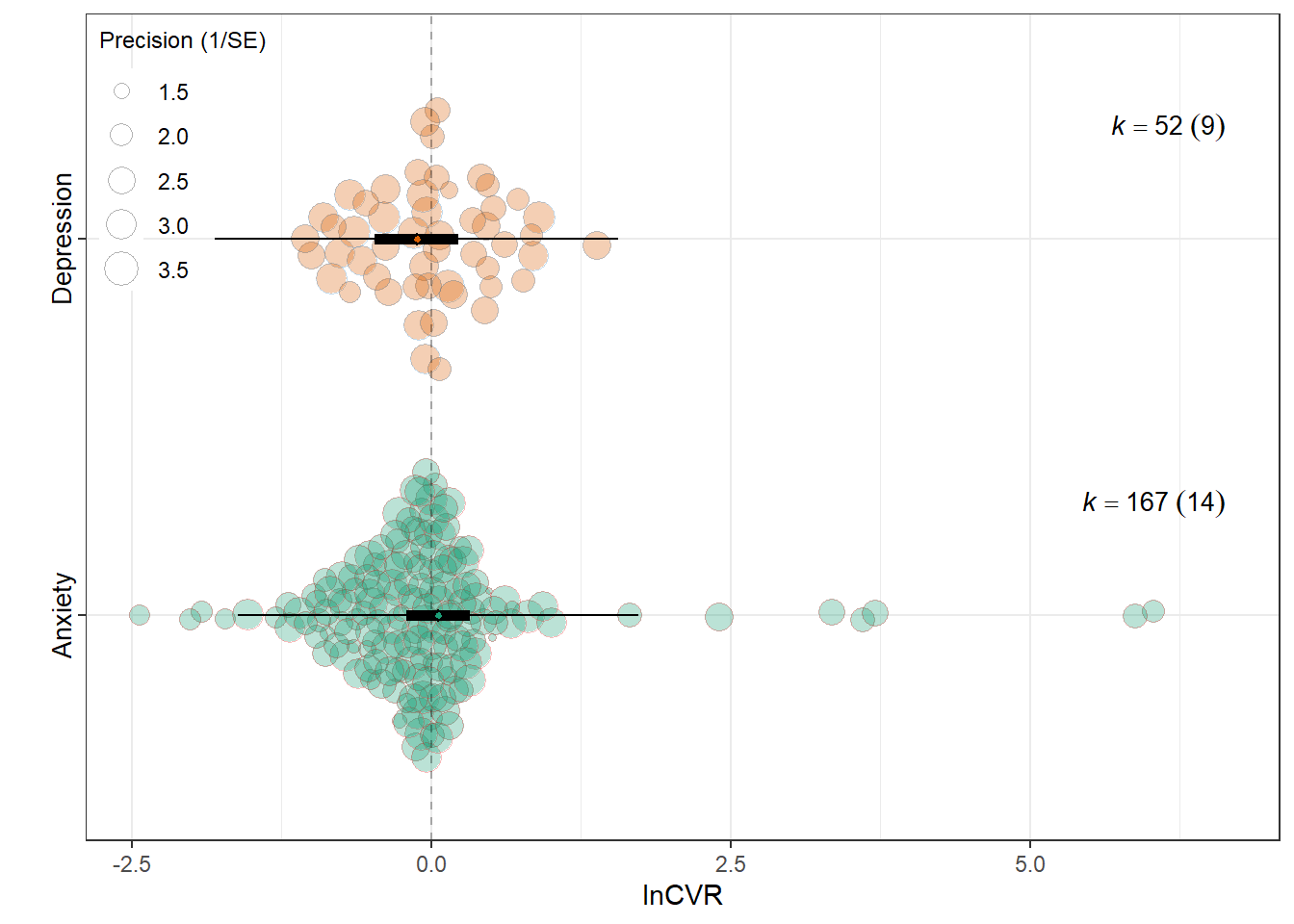
Behavioral assay
MOD <- "Assay_type"
MODS <- "~ Assay_type - 1"
assay_labels <- c(
"Elevated Plus Maze" = "EPM",
"Open Field Test" = "OFT",
"Light-Dark Box" = "LDB",
"Forced Swim Test (FST)" = "FST",
"Tail Suspension Test (TST)" = "TST",
"Sucrose Preference Test (SPT)" = "SPT"
)
# 1) Filter each effect-size dataset (and subset the VCV accordingly)
f_rr <- filter_low_k_single(db_rr, VCV_rr, MOD, min_k = 5)
f_vr <- filter_low_k_single(db_vr, VCV_vr, MOD, min_k = 5)
f_cvr <- filter_low_k_single(db_cvr, VCV_cvr, MOD, min_k = 5)
# 2) Fit the three unimoderator models (same moderator, different yi/vi)
res_rr <- fit_unimod(f_rr$data, f_rr$V, "lnRR", MODS, MOD, expression(lnRR))
res_vr <- fit_unimod(f_vr$data, f_vr$V, "lnVR", MODS, MOD, expression(lnVR))
res_cvr <- fit_unimod(f_cvr$data, f_cvr$V, "lnCVR", MODS, MOD, expression(lnCVR))
# 2b) Relabel x-axis in the orchard plots (so EPM/OFT/etc. show)
res_rr$plot <- res_rr$plot + scale_x_discrete(labels = assay_labels)
res_vr$plot <- res_vr$plot + scale_x_discrete(labels = assay_labels)
res_cvr$plot <- res_cvr$plot + scale_x_discrete(labels = assay_labels)
# 3) Tukey contrasts (use the filtered data + matched VCV for each dataset)
tuk_rr <- maybe_tukey(dat = f_rr$data, V = f_rr$V, yi = "lnRR", moderator = MOD)
tuk_vr <- maybe_tukey(dat = f_vr$data, V = f_vr$V, yi = "lnVR", moderator = MOD)
tuk_cvr <- maybe_tukey(dat = f_cvr$data, V = f_cvr$V, yi = "lnCVR", moderator = MOD)lnRR
summary(res_rr$model)
Multivariate Meta-Analysis Model (k = 289; method: REML)
logLik Deviance AIC BIC AICc
-227.2656 454.5311 472.5311 505.3401 473.1904
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0311 0.1763 20 no Study_ID
sigma^2.2 0.2267 0.4762 289 no ES_ID
sigma^2.3 0.0043 0.0654 6 no Strain
Test for Residual Heterogeneity:
QE(df = 283) = 5482.7134, p-val < .0001
Test of Moderators (coefficients 1:6):
F(df1 = 6, df2 = 283) = 3.8229, p-val = 0.0011
Model Results:
estimate se tval df pval
Assay_typeElevated Plus Maze -0.2884 0.0805 -3.5807 283 0.0004
Assay_typeForced Swim Test (FST) -0.3122 0.1127 -2.7713 283 0.0060
Assay_typeLight-Dark Box -0.5634 0.1518 -3.7114 283 0.0002
Assay_typeOpen Field Test -0.0699 0.0844 -0.8280 283 0.4084
Assay_typeSucrose Preference Test (SPT) -0.0715 0.1574 -0.4542 283 0.6500
Assay_typeTail Suspension Test (TST) -0.0926 0.2025 -0.4571 283 0.6480
ci.lb ci.ub
Assay_typeElevated Plus Maze -0.4469 -0.1298 ***
Assay_typeForced Swim Test (FST) -0.5339 -0.0905 **
Assay_typeLight-Dark Box -0.8622 -0.2646 ***
Assay_typeOpen Field Test -0.2360 0.0963
Assay_typeSucrose Preference Test (SPT) -0.3813 0.2383
Assay_typeTail Suspension Test (TST) -0.4912 0.3061
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_rr$r2 R2_marginal R2_conditional
0.0709 0.1963 res_rr$plot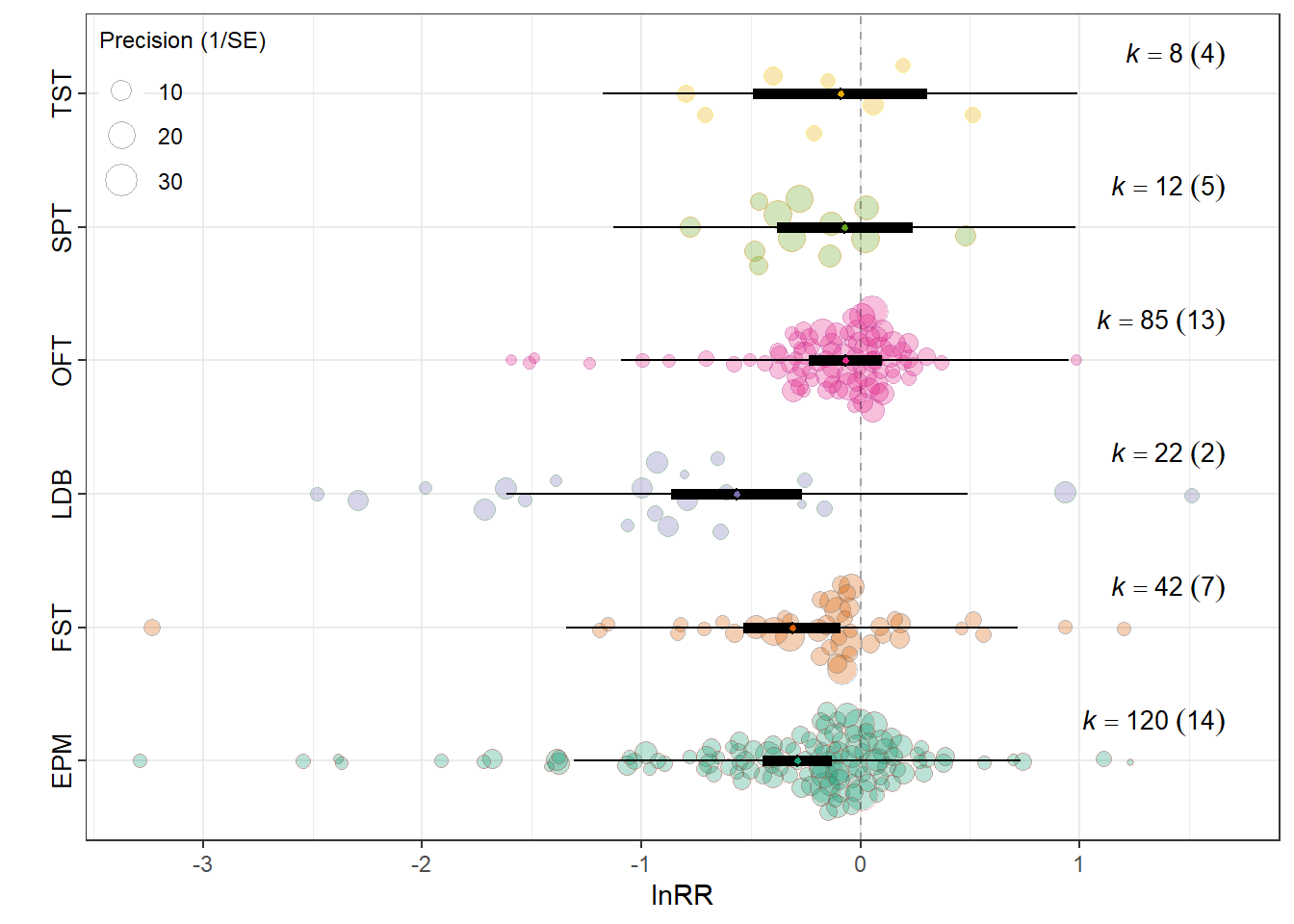
tuk_rr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 -0.023825 0.112008 -0.213 0.83155
3 - 1 == 0 -0.275002 0.136691 -2.012 0.04424 *
4 - 1 == 0 0.218483 0.083046 2.631 0.00852 **
5 - 1 == 0 0.216889 0.162745 1.333 0.18263
6 - 1 == 0 0.195807 0.207107 0.945 0.34443
3 - 2 == 0 -0.251177 0.172914 -1.453 0.14633
4 - 2 == 0 0.242308 0.119213 2.033 0.04210 *
5 - 2 == 0 0.240714 0.179481 1.341 0.17987
6 - 2 == 0 0.219632 0.221942 0.990 0.32237
4 - 3 == 0 0.493485 0.153496 3.215 0.00130 **
5 - 3 == 0 0.491891 0.208187 2.363 0.01814 *
6 - 3 == 0 0.470809 0.244179 1.928 0.05384 .
5 - 4 == 0 -0.001594 0.163517 -0.010 0.99222
6 - 4 == 0 -0.022676 0.206881 -0.110 0.91272
6 - 5 == 0 -0.021082 0.243796 -0.086 0.93109
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Adjusted p values reported -- none method)lnVR
summary(res_vr$model)
Multivariate Meta-Analysis Model (k = 213; method: REML)
logLik Deviance AIC BIC AICc
-321.7773 643.5546 659.5546 686.2549 660.2783
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.1545 0.3931 16 no Study_ID
sigma^2.2 1.1373 1.0665 213 no ES_ID
sigma^2.3 0.0000 0.0001 6 no Strain
Test for Residual Heterogeneity:
QE(df = 208) = 3885.5939, p-val < .0001
Test of Moderators (coefficients 1:5):
F(df1 = 5, df2 = 208) = 4.4278, p-val = 0.0007
Model Results:
estimate se tval df pval
Assay_typeElevated Plus Maze 0.7333 0.1947 3.7653 208 0.0002
Assay_typeForced Swim Test (FST) -0.1735 0.2472 -0.7019 208 0.4835
Assay_typeLight-Dark Box 0.1806 0.3190 0.5659 208 0.5720
Assay_typeOpen Field Test -0.0219 0.1924 -0.1138 208 0.9095
Assay_typeTail Suspension Test (TST) -0.5177 0.4369 -1.1850 208 0.2374
ci.lb ci.ub
Assay_typeElevated Plus Maze 0.3493 1.1172 ***
Assay_typeForced Swim Test (FST) -0.6609 0.3139
Assay_typeLight-Dark Box -0.4484 0.8095
Assay_typeOpen Field Test -0.4011 0.3573
Assay_typeTail Suspension Test (TST) -1.3789 0.3436
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_vr$r2 R2_marginal R2_conditional
0.1119 0.2181 res_vr$plot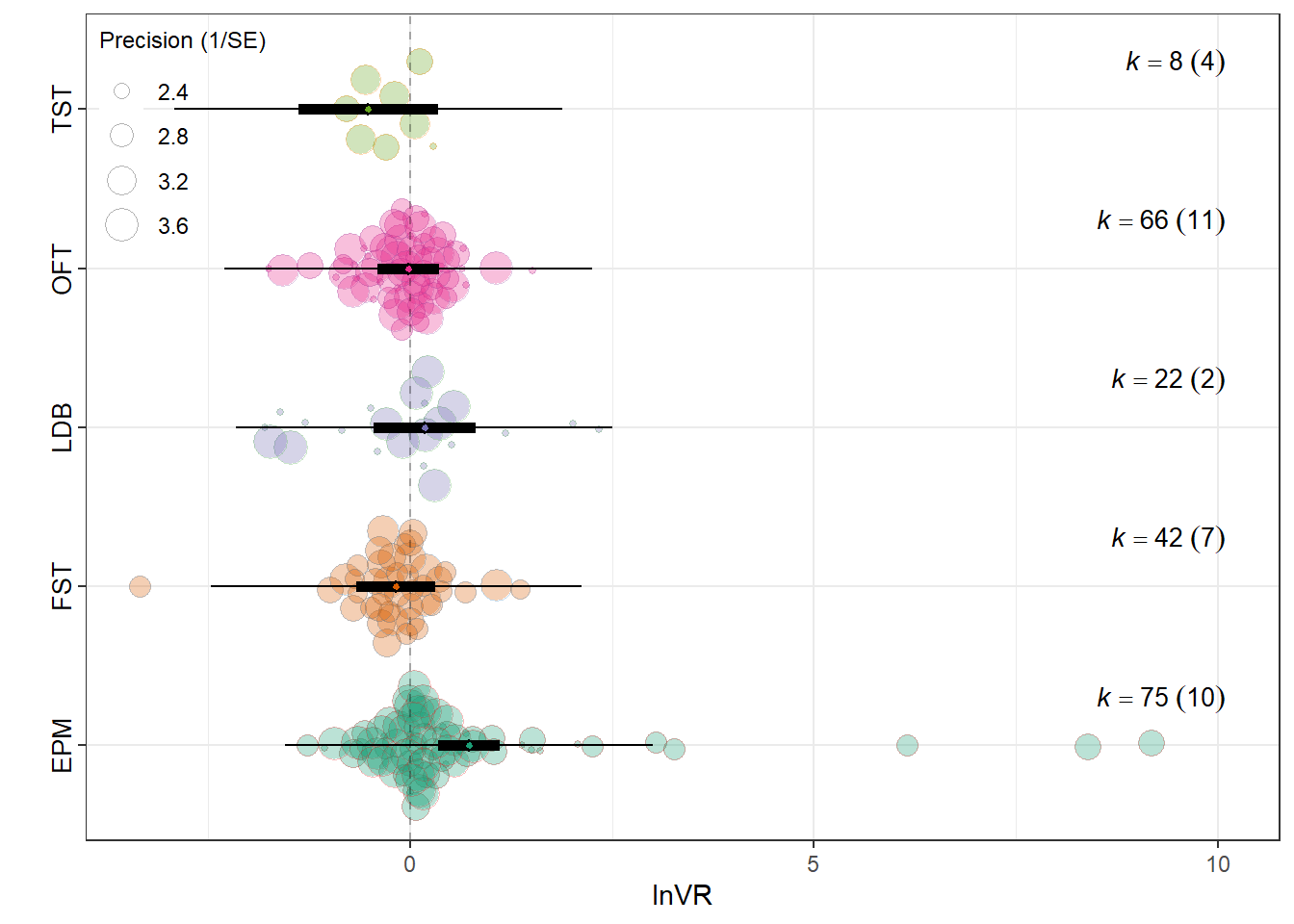
tuk_vr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 -0.9068 0.2687 -3.375 0.000739 ***
3 - 1 == 0 -0.5527 0.2909 -1.900 0.057391 .
4 - 1 == 0 -0.7552 0.2241 -3.369 0.000754 ***
5 - 1 == 0 -1.2510 0.4611 -2.713 0.006670 **
3 - 2 == 0 0.3541 0.3775 0.938 0.348283
4 - 2 == 0 0.1516 0.2743 0.553 0.580379
5 - 2 == 0 -0.3442 0.4877 -0.706 0.480382
4 - 3 == 0 -0.2024 0.3416 -0.593 0.553443
5 - 3 == 0 -0.6982 0.5292 -1.319 0.187032
5 - 4 == 0 -0.4958 0.4573 -1.084 0.278306
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Adjusted p values reported -- none method)lnCVR
summary(res_cvr$model)
Multivariate Meta-Analysis Model (k = 213; method: REML)
logLik Deviance AIC BIC AICc
-267.8978 535.7956 551.7956 578.4959 552.5192
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.1244 0.3526 16 no Study_ID
sigma^2.2 0.5920 0.7694 213 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 208) = 1504.7562, p-val < .0001
Test of Moderators (coefficients 1:5):
F(df1 = 5, df2 = 208) = 1.1484, p-val = 0.3360
Model Results:
estimate se tval df pval
Assay_typeElevated Plus Maze 0.2586 0.1667 1.5511 208 0.1224
Assay_typeForced Swim Test (FST) -0.0550 0.2029 -0.2709 208 0.7867
Assay_typeLight-Dark Box 0.1576 0.2660 0.5924 208 0.5543
Assay_typeOpen Field Test -0.1324 0.1622 -0.8164 208 0.4152
Assay_typeTail Suspension Test (TST) -0.1191 0.3430 -0.3473 208 0.7287
ci.lb ci.ub
Assay_typeElevated Plus Maze -0.0701 0.5872
Assay_typeForced Swim Test (FST) -0.4550 0.3451
Assay_typeLight-Dark Box -0.3669 0.6820
Assay_typeOpen Field Test -0.4521 0.1873
Assay_typeTail Suspension Test (TST) -0.7953 0.5571
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_cvr$r2 R2_marginal R2_conditional
0.0405 0.2070 res_cvr$plot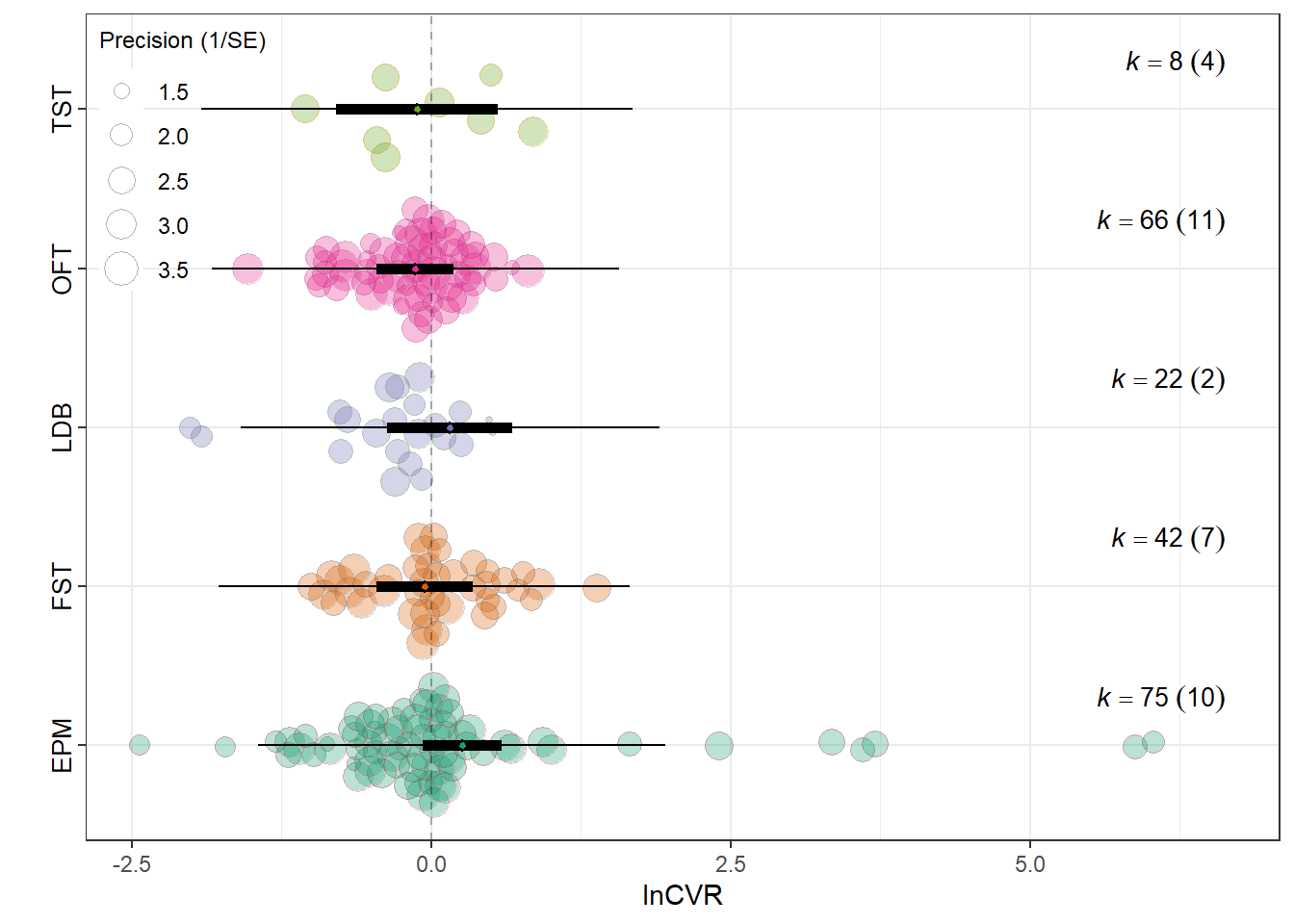
tuk_cvr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 -0.31354 0.20741 -1.512 0.1306
3 - 1 == 0 -0.10099 0.22731 -0.444 0.6568
4 - 1 == 0 -0.39097 0.17392 -2.248 0.0246 *
5 - 1 == 0 -0.37769 0.35917 -1.052 0.2930
3 - 2 == 0 0.21255 0.29885 0.711 0.4769
4 - 2 == 0 -0.07743 0.21225 -0.365 0.7153
5 - 2 == 0 -0.06415 0.37941 -0.169 0.8657
4 - 3 == 0 -0.28998 0.27232 -1.065 0.2869
5 - 3 == 0 -0.27670 0.41692 -0.664 0.5069
5 - 4 == 0 0.01329 0.35474 0.037 0.9701
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Adjusted p values reported -- none method)Behavioral mechanism
MOD <- "Induced behaviour"
MODS <- "~ `Induced behaviour`"
f_rr <- filter_low_k_single(db_rr, VCV_rr, MOD, min_k = 5)
f_vr <- filter_low_k_single(db_vr, VCV_vr, MOD, min_k = 5)
f_cvr <- filter_low_k_single(db_cvr, VCV_cvr, MOD, min_k = 5)
res_rr <- fit_unimod(f_rr$data, f_rr$V, "lnRR", MODS, MOD, expression(lnRR))
res_vr <- fit_unimod(f_vr$data, f_vr$V, "lnVR", MODS, MOD, expression(lnVR))
res_cvr <- fit_unimod(f_cvr$data, f_cvr$V, "lnCVR", MODS, MOD, expression(lnCVR))lnRR
summary(res_rr$model)
Multivariate Meta-Analysis Model (k = 298; method: REML)
logLik Deviance AIC BIC AICc
-238.1126 476.2252 486.2252 504.6770 486.4321
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0484 0.2199 20 no Study_ID
sigma^2.2 0.2228 0.4720 298 no ES_ID
sigma^2.3 0.0000 0.0001 6 no Strain
Test for Residual Heterogeneity:
QE(df = 296) = 5461.1248, p-val < .0001
Test of Moderators (coefficient 2):
F(df1 = 1, df2 = 296) = 8.4650, p-val = 0.0039
Model Results:
estimate se tval df pval ci.lb
intrcpt -0.4579 0.1072 -4.2731 296 <.0001 -0.6688
`Induced behaviour`Innate 0.3222 0.1107 2.9095 296 0.0039 0.1043
ci.ub
intrcpt -0.2470 ***
`Induced behaviour`Innate 0.5401 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_rr$r2 R2_marginal R2_conditional
0.0524 0.2214 res_rr$plot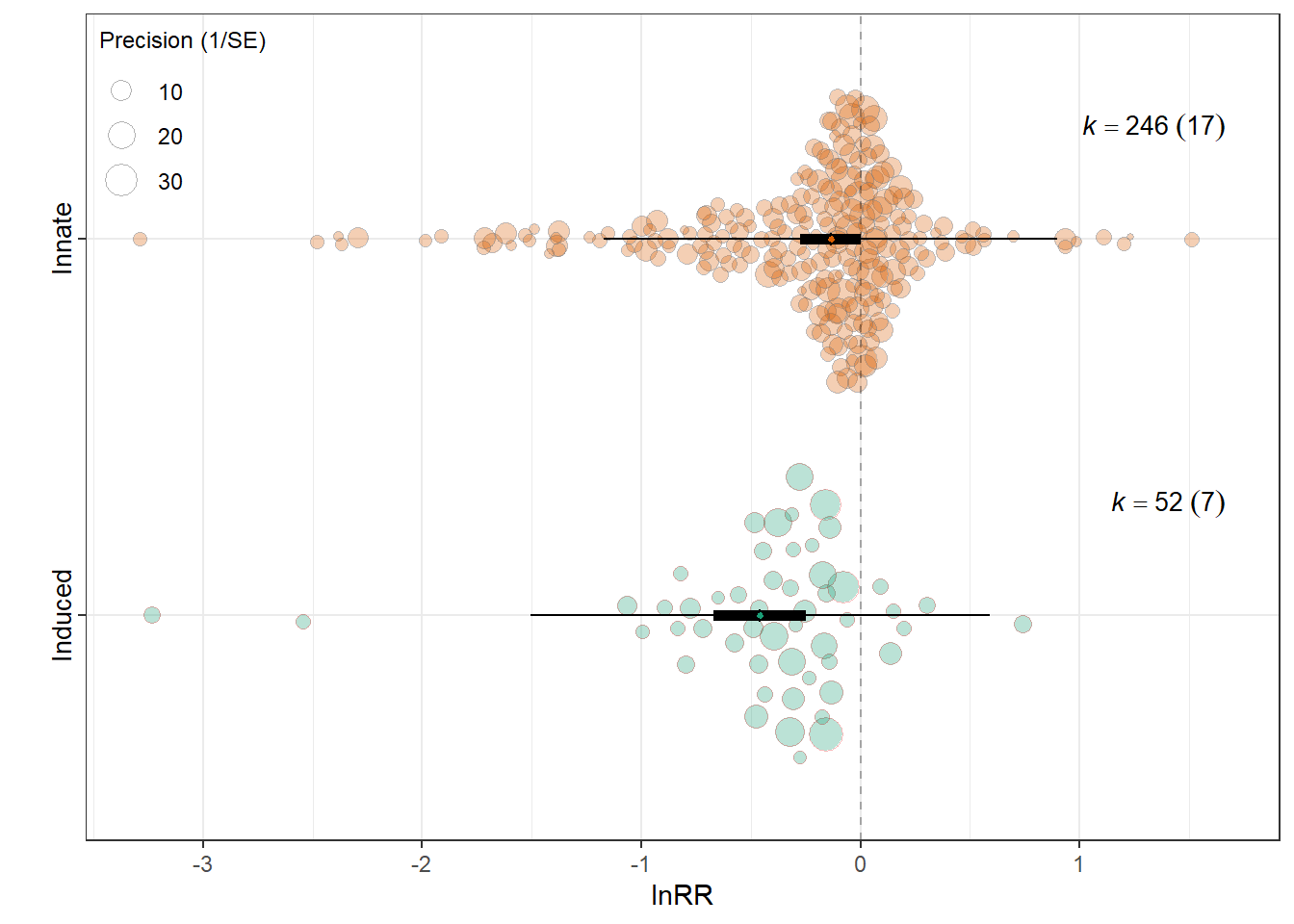
lnVR
summary(res_vr$model)
Multivariate Meta-Analysis Model (k = 222; method: REML)
logLik Deviance AIC BIC AICc
-346.5856 693.1713 703.1713 720.1394 703.4517
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0926 0.3043 16 no Study_ID
sigma^2.2 1.2261 1.1073 222 no ES_ID
sigma^2.3 0.0000 0.0001 6 no Strain
Test for Residual Heterogeneity:
QE(df = 220) = 4389.9716, p-val < .0001
Test of Moderators (coefficient 2):
F(df1 = 1, df2 = 220) = 0.2942, p-val = 0.5881
Model Results:
estimate se tval df pval ci.lb
intrcpt 0.2494 0.2478 1.0064 220 0.3153 -0.2390
`Induced behaviour`Innate -0.1461 0.2694 -0.5424 220 0.5881 -0.6771
ci.ub
intrcpt 0.7378
`Induced behaviour`Innate 0.3848
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_vr$r2 R2_marginal R2_conditional
0.0022 0.0723 res_vr$plot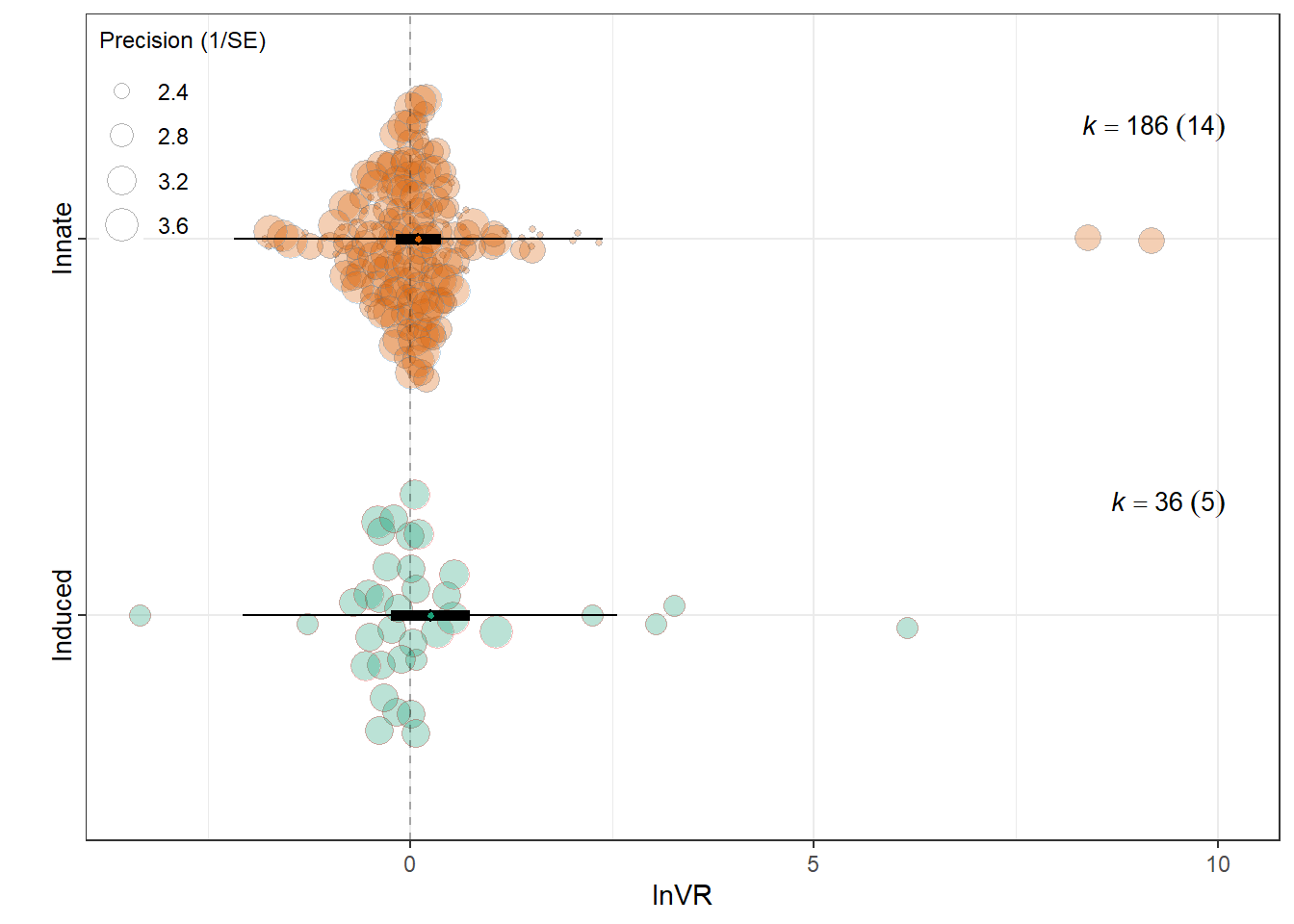
lnCVR
summary(res_cvr$model)
Multivariate Meta-Analysis Model (k = 222; method: REML)
logLik Deviance AIC BIC AICc
-282.2780 564.5560 574.5560 591.5241 574.8364
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0766 0.2768 16 no Study_ID
sigma^2.2 0.5953 0.7715 222 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 220) = 1588.3881, p-val < .0001
Test of Moderators (coefficient 2):
F(df1 = 1, df2 = 220) = 0.5346, p-val = 0.4655
Model Results:
estimate se tval df pval ci.lb
intrcpt 0.1362 0.2021 0.6741 220 0.5010 -0.2621
`Induced behaviour`Innate -0.1563 0.2138 -0.7311 220 0.4655 -0.5777
ci.ub
intrcpt 0.5345
`Induced behaviour`Innate 0.2651
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_cvr$r2 R2_marginal R2_conditional
0.0049 0.1184 res_cvr$plot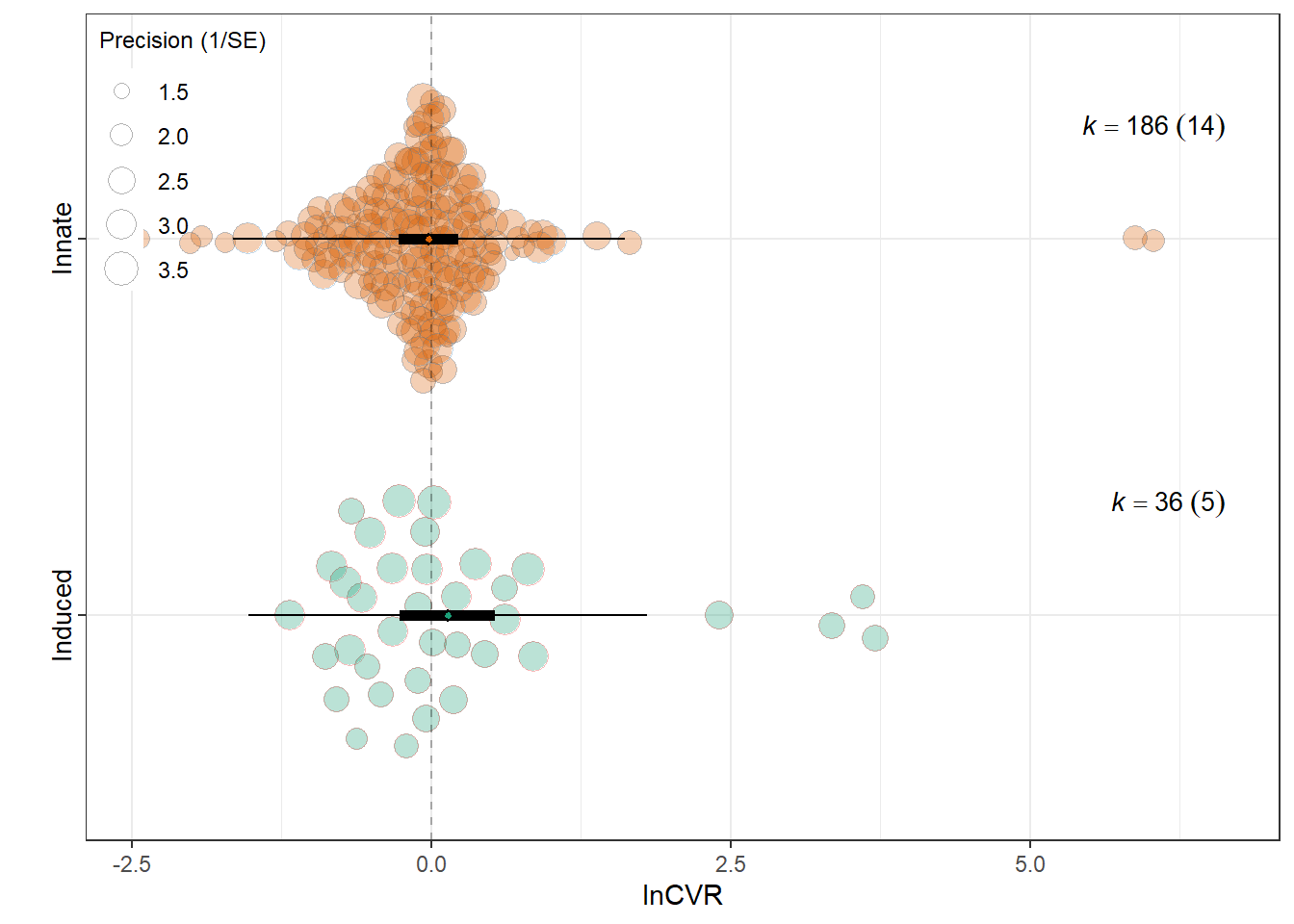
Lifestage of exposure
MOD <- "Lifestage_exposure"
MODS <- "~ Lifestage_exposure -1"
lifestage_labels <- c(
"Adolescent" = "Adol.",
"Adult" = "Adult",
"Juvenile" = "Juv.",
"Young adult" = "Y. adult",
"Mixed" = "Mixed",
"Unclear" = "Unclear"
)
# 1) Filter each effect-size dataset (and subset the VCV accordingly)
f_rr <- filter_low_k_single(db_rr, VCV_rr, MOD, min_k = 5)
f_vr <- filter_low_k_single(db_vr, VCV_vr, MOD, min_k = 5)
f_cvr <- filter_low_k_single(db_cvr, VCV_cvr, MOD, min_k = 5)
# 2) Fit the three unimoderator models (same moderator, different yi/vi)
res_rr <- fit_unimod(f_rr$data, f_rr$V, "lnRR", MODS, MOD, expression(lnRR))
res_vr <- fit_unimod(f_vr$data, f_vr$V, "lnVR", MODS, MOD, expression(lnVR))
res_cvr <- fit_unimod(f_cvr$data, f_cvr$V, "lnCVR", MODS, MOD, expression(lnCVR))
# 2b) Relabel x-axis in the orchard plots (so EPM/OFT/etc. show)
res_rr$plot <- res_rr$plot + scale_x_discrete(labels = lifestage_labels)
res_vr$plot <- res_vr$plot + scale_x_discrete(labels = lifestage_labels)
res_cvr$plot <- res_cvr$plot + scale_x_discrete(labels = lifestage_labels)
# 3) Tukey contrasts (use the filtered data + matched VCV for each dataset)
tuk_rr <- maybe_tukey(dat = f_rr$data, V = f_rr$V, yi = "lnRR", moderator = MOD)
tuk_vr <- maybe_tukey(dat = f_vr$data, V = f_vr$V, yi = "lnVR", moderator = MOD)
tuk_cvr <- maybe_tukey(dat = f_cvr$data, V = f_cvr$V, yi = "lnCVR", moderator = MOD)lnRR
summary(res_rr$model)
Multivariate Meta-Analysis Model (k = 298; method: REML)
logLik Deviance AIC BIC AICc
-238.0800 476.1601 494.1601 527.2508 494.7983
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0390 0.1974 20 no Study_ID
sigma^2.2 0.2333 0.4830 298 no ES_ID
sigma^2.3 0.0045 0.0670 6 no Strain
Test for Residual Heterogeneity:
QE(df = 292) = 5634.2828, p-val < .0001
Test of Moderators (coefficients 1:6):
F(df1 = 6, df2 = 292) = 2.0555, p-val = 0.0585
Model Results:
estimate se tval df pval ci.lb
Lifestage_exposureAdolescent -0.2269 0.1630 -1.3921 292 0.1650 -0.5476
Lifestage_exposureAdult -0.1332 0.1448 -0.9199 292 0.3584 -0.4182
Lifestage_exposureJuvenile -0.1303 0.1946 -0.6698 292 0.5035 -0.5134
Lifestage_exposureMixed -0.0596 0.1184 -0.5034 292 0.6151 -0.2927
Lifestage_exposureUnclear -0.3490 0.1678 -2.0800 292 0.0384 -0.6793
Lifestage_exposureYoung adult -0.3083 0.0909 -3.3914 292 0.0008 -0.4873
ci.ub
Lifestage_exposureAdolescent 0.0939
Lifestage_exposureAdult 0.1518
Lifestage_exposureJuvenile 0.2527
Lifestage_exposureMixed 0.1735
Lifestage_exposureUnclear -0.0188 *
Lifestage_exposureYoung adult -0.1294 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_rr$r2 R2_marginal R2_conditional
0.0415 0.1920 res_rr$plot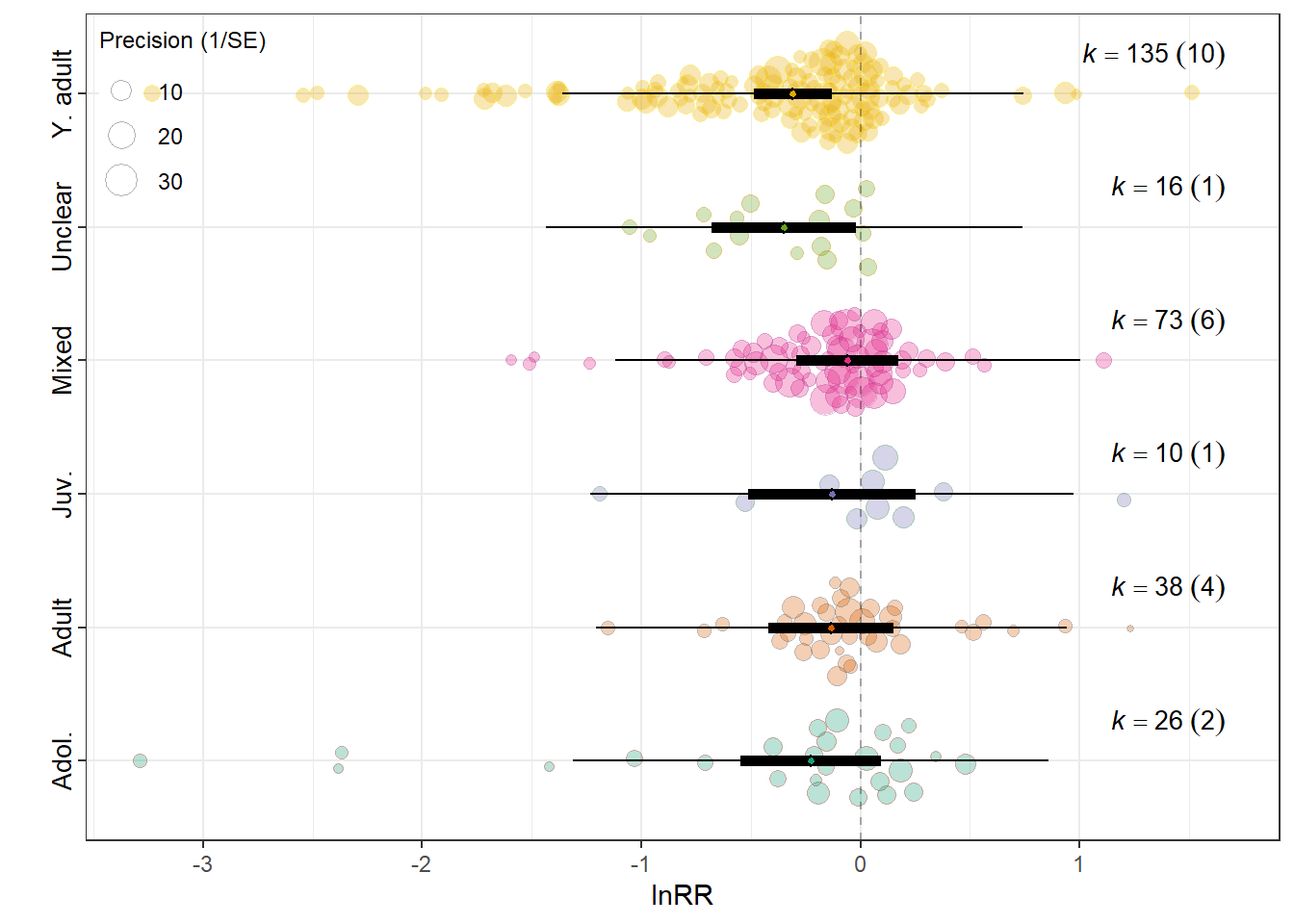
tuk_rr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 0.093666 0.186134 0.503 0.6148
3 - 1 == 0 0.096523 0.208702 0.462 0.6437
4 - 1 == 0 0.167255 0.197336 0.848 0.3967
5 - 1 == 0 -0.122164 0.220161 -0.555 0.5790
6 - 1 == 0 -0.081458 0.166351 -0.490 0.6244
3 - 2 == 0 0.002857 0.206027 0.014 0.9889
4 - 2 == 0 0.073588 0.180845 0.407 0.6841
5 - 2 == 0 -0.215830 0.209449 -1.030 0.3028
6 - 2 == 0 -0.175125 0.152363 -1.149 0.2504
4 - 3 == 0 0.070731 0.223106 0.317 0.7512
5 - 3 == 0 -0.218687 0.241529 -0.905 0.3652
6 - 3 == 0 -0.177982 0.193208 -0.921 0.3569
5 - 4 == 0 -0.289418 0.201759 -1.434 0.1514
6 - 4 == 0 -0.248713 0.143526 -1.733 0.0831 .
6 - 5 == 0 0.040705 0.148271 0.275 0.7837
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Adjusted p values reported -- none method)lnVR
summary(res_vr$model)
Multivariate Meta-Analysis Model (k = 222; method: REML)
logLik Deviance AIC BIC AICc
-339.3525 678.7050 696.7050 727.0825 697.5788
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0263 0.1621 16 no Study_ID
sigma^2.2 1.2491 1.1176 222 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 216) = 4359.9557, p-val < .0001
Test of Moderators (coefficients 1:6):
F(df1 = 6, df2 = 216) = 1.1160, p-val = 0.3539
Model Results:
estimate se tval df pval ci.lb
Lifestage_exposureAdolescent 0.7391 0.3236 2.2844 216 0.0233 0.1014
Lifestage_exposureAdult -0.0508 0.2667 -0.1903 216 0.8492 -0.5764
Lifestage_exposureJuvenile 0.0347 0.5057 0.0685 216 0.9454 -0.9621
Lifestage_exposureMixed -0.1120 0.2389 -0.4687 216 0.6398 -0.5829
Lifestage_exposureUnclear 0.1922 0.3773 0.5094 216 0.6110 -0.5515
Lifestage_exposureYoung adult 0.1715 0.1561 1.0988 216 0.2731 -0.1361
ci.ub
Lifestage_exposureAdolescent 1.3768 *
Lifestage_exposureAdult 0.4749
Lifestage_exposureJuvenile 1.0315
Lifestage_exposureMixed 0.3589
Lifestage_exposureUnclear 0.9359
Lifestage_exposureYoung adult 0.4791
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_vr$r2 R2_marginal R2_conditional
0.0388 0.0586 res_vr$plot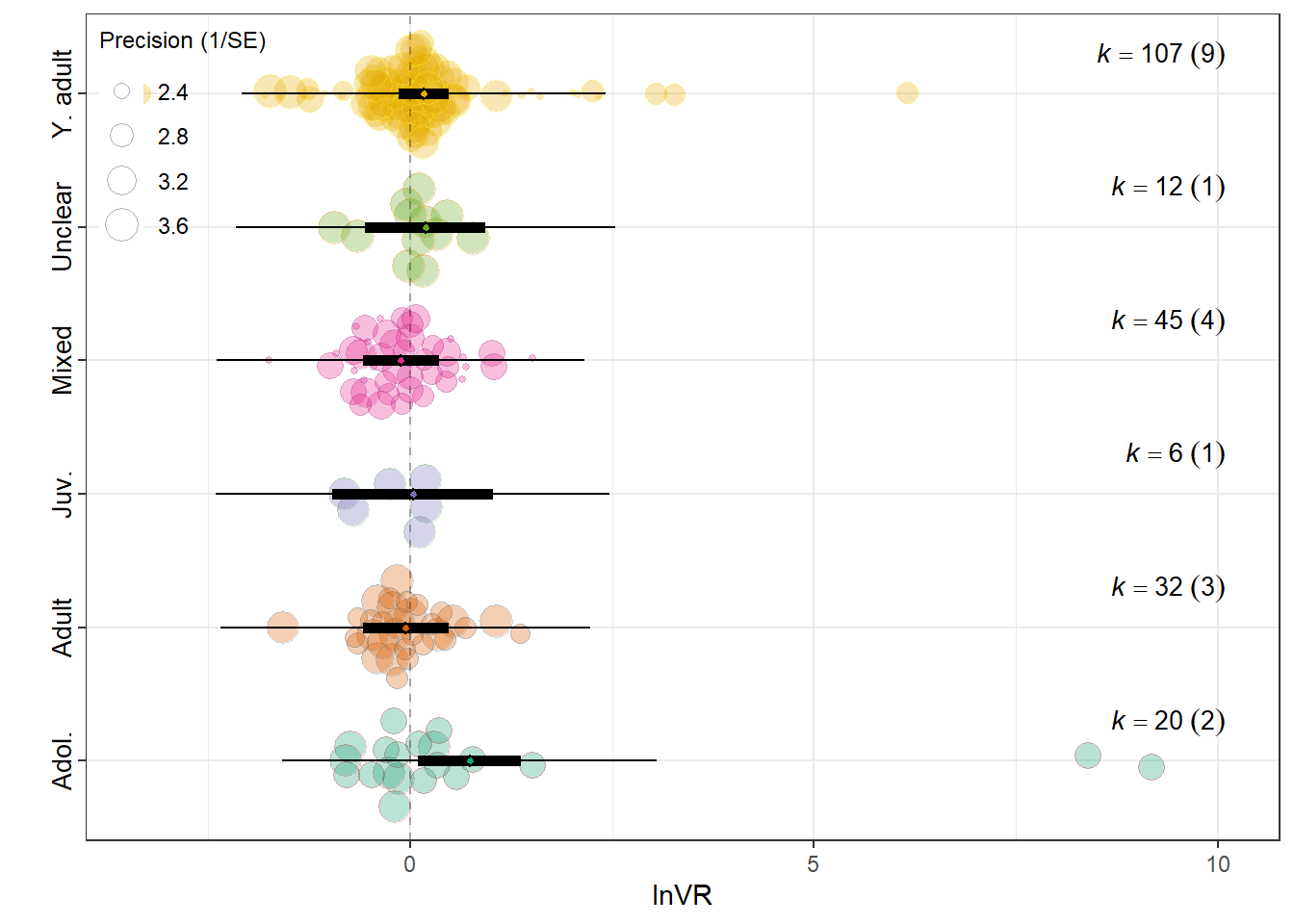
tuk_vr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 -0.78987 0.40549 -1.948 0.0514 .
3 - 1 == 0 -0.70446 0.56920 -1.238 0.2159
4 - 1 == 0 -0.85110 0.40220 -2.116 0.0343 *
5 - 1 == 0 -0.54692 0.49456 -1.106 0.2688
6 - 1 == 0 -0.56763 0.35356 -1.605 0.1084
3 - 2 == 0 0.08541 0.54910 0.156 0.8764
4 - 2 == 0 -0.06122 0.35804 -0.171 0.8642
5 - 2 == 0 0.24296 0.46018 0.528 0.5975
6 - 2 == 0 0.22225 0.30441 0.730 0.4653
4 - 3 == 0 -0.14663 0.55932 -0.262 0.7932
5 - 3 == 0 0.15754 0.62662 0.251 0.8015
6 - 3 == 0 0.13683 0.52068 0.263 0.7927
5 - 4 == 0 0.30418 0.44658 0.681 0.4958
6 - 4 == 0 0.28347 0.28538 0.993 0.3206
6 - 5 == 0 -0.02071 0.37038 -0.056 0.9554
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Adjusted p values reported -- none method)lnCVR
summary(res_cvr$model)
Multivariate Meta-Analysis Model (k = 222; method: REML)
logLik Deviance AIC BIC AICc
-275.4287 550.8573 568.8573 599.2348 569.7311
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0914 0.3023 16 no Study_ID
sigma^2.2 0.5881 0.7669 222 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 216) = 1529.9807, p-val < .0001
Test of Moderators (coefficients 1:6):
F(df1 = 6, df2 = 216) = 0.8899, p-val = 0.5031
Model Results:
estimate se tval df pval ci.lb
Lifestage_exposureAdolescent 0.2796 0.3072 0.9103 216 0.3637 -0.3258
Lifestage_exposureAdult -0.1264 0.2565 -0.4929 216 0.6226 -0.6319
Lifestage_exposureJuvenile -0.0182 0.3922 -0.0464 216 0.9631 -0.7913
Lifestage_exposureMixed -0.2921 0.2504 -1.1668 216 0.2446 -0.7856
Lifestage_exposureUnclear -0.1622 0.3034 -0.5346 216 0.5935 -0.7601
Lifestage_exposureYoung adult 0.1846 0.1625 1.1359 216 0.2573 -0.1357
ci.ub
Lifestage_exposureAdolescent 0.8851
Lifestage_exposureAdult 0.3791
Lifestage_exposureJuvenile 0.7549
Lifestage_exposureMixed 0.2014
Lifestage_exposureUnclear 0.4358
Lifestage_exposureYoung adult 0.5048
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_cvr$r2 R2_marginal R2_conditional
0.0607 0.1871 res_cvr$plot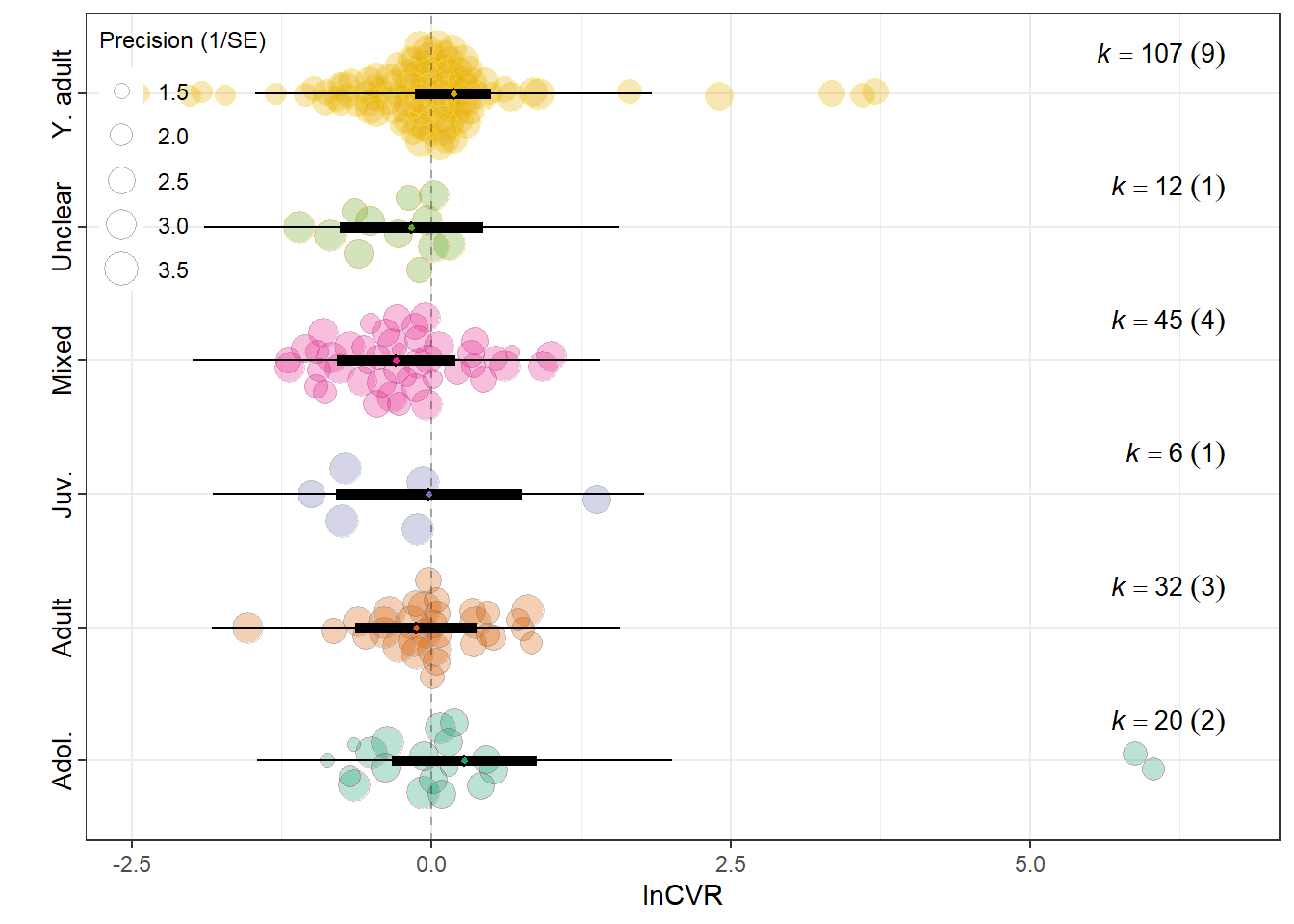
tuk_cvr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 -0.40605 0.35824 -1.133 0.257
3 - 1 == 0 -0.29783 0.43007 -0.693 0.489
4 - 1 == 0 -0.57177 0.39630 -1.443 0.149
5 - 1 == 0 -0.44183 0.41766 -1.058 0.290
6 - 1 == 0 -0.09509 0.32661 -0.291 0.771
3 - 2 == 0 0.10822 0.41552 0.260 0.795
4 - 2 == 0 -0.16572 0.35841 -0.462 0.644
5 - 2 == 0 -0.03579 0.38591 -0.093 0.926
6 - 2 == 0 0.31096 0.28585 1.088 0.277
4 - 3 == 0 -0.27394 0.46532 -0.589 0.556
5 - 3 == 0 -0.14400 0.47772 -0.301 0.763
6 - 3 == 0 0.20274 0.39930 0.508 0.612
5 - 4 == 0 0.12994 0.39335 0.330 0.741
6 - 4 == 0 0.47668 0.29847 1.597 0.110
6 - 5 == 0 0.34674 0.27131 1.278 0.201
(Adjusted p values reported -- none method)Control conditions
MOD <- "Control_conditions"
MODS <- "~ `Control_conditions`"
f_rr <- filter_low_k_single(db_rr, VCV_rr, MOD, min_k = 5)
f_vr <- filter_low_k_single(db_vr, VCV_vr, MOD, min_k = 5)
f_cvr <- filter_low_k_single(db_cvr, VCV_cvr, MOD, min_k = 5)
res_rr <- fit_unimod(f_rr$data, f_rr$V, "lnRR", MODS, MOD, expression(lnRR))
res_vr <- fit_unimod(f_vr$data, f_vr$V, "lnVR", MODS, MOD, expression(lnVR))
res_cvr <- fit_unimod(f_cvr$data, f_cvr$V, "lnCVR", MODS, MOD, expression(lnCVR))lnRR
summary(res_rr$model)
Multivariate Meta-Analysis Model (k = 298; method: REML)
logLik Deviance AIC BIC AICc
-239.1884 478.3768 488.3768 506.8286 488.5837
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0454 0.2131 20 no Study_ID
sigma^2.2 0.2260 0.4754 298 no ES_ID
sigma^2.3 0.0000 0.0001 6 no Strain
Test for Residual Heterogeneity:
QE(df = 296) = 5396.0343, p-val < .0001
Test of Moderators (coefficient 2):
F(df1 = 1, df2 = 296) = 6.5356, p-val = 0.0111
Model Results:
estimate se tval df pval ci.lb
intrcpt -0.1871 0.0650 -2.8775 296 0.0043 -0.3151
Control_conditionsWhite noise -0.2618 0.1024 -2.5565 296 0.0111 -0.4634
ci.ub
intrcpt -0.0591 **
Control_conditionsWhite noise -0.0603 *
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_rr$r2 R2_marginal R2_conditional
0.0315 0.1936 res_rr$plot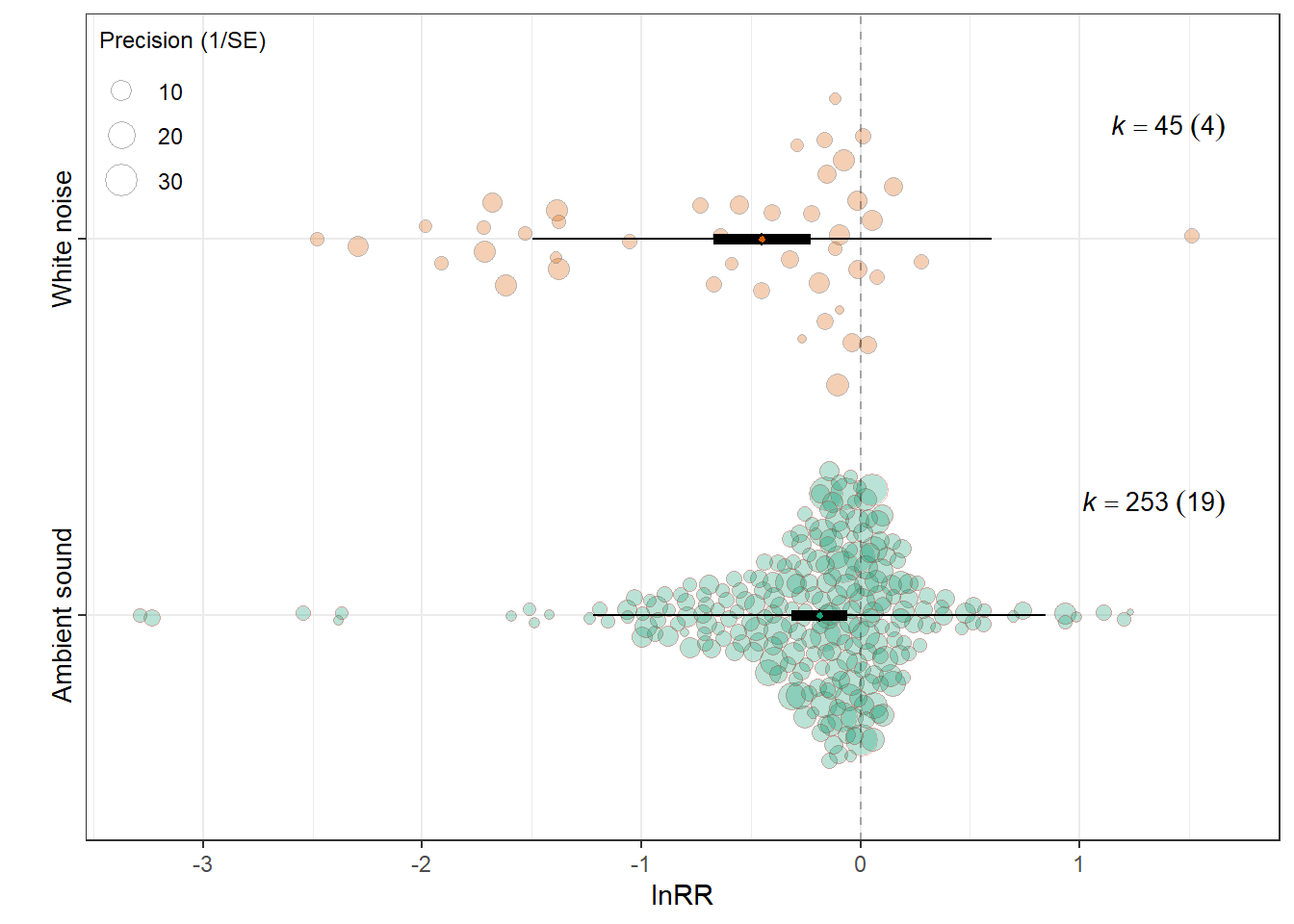
lnVR
summary(res_vr$model)
Multivariate Meta-Analysis Model (k = 222; method: REML)
logLik Deviance AIC BIC AICc
-346.6305 693.2609 703.2609 720.2291 703.5413
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0857 0.2927 16 no Study_ID
sigma^2.2 1.2273 1.1078 222 no ES_ID
sigma^2.3 0.0000 0.0001 6 no Strain
Test for Residual Heterogeneity:
QE(df = 220) = 4380.6933, p-val < .0001
Test of Moderators (coefficient 2):
F(df1 = 1, df2 = 220) = 0.3657, p-val = 0.5460
Model Results:
estimate se tval df pval ci.lb
intrcpt 0.1497 0.1311 1.1415 220 0.2549 -0.1087
Control_conditionsWhite noise -0.1533 0.2535 -0.6047 220 0.5460 -0.6530
ci.ub
intrcpt 0.4081
Control_conditionsWhite noise 0.3463
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_vr$r2 R2_marginal R2_conditional
0.0023 0.0674 res_vr$plot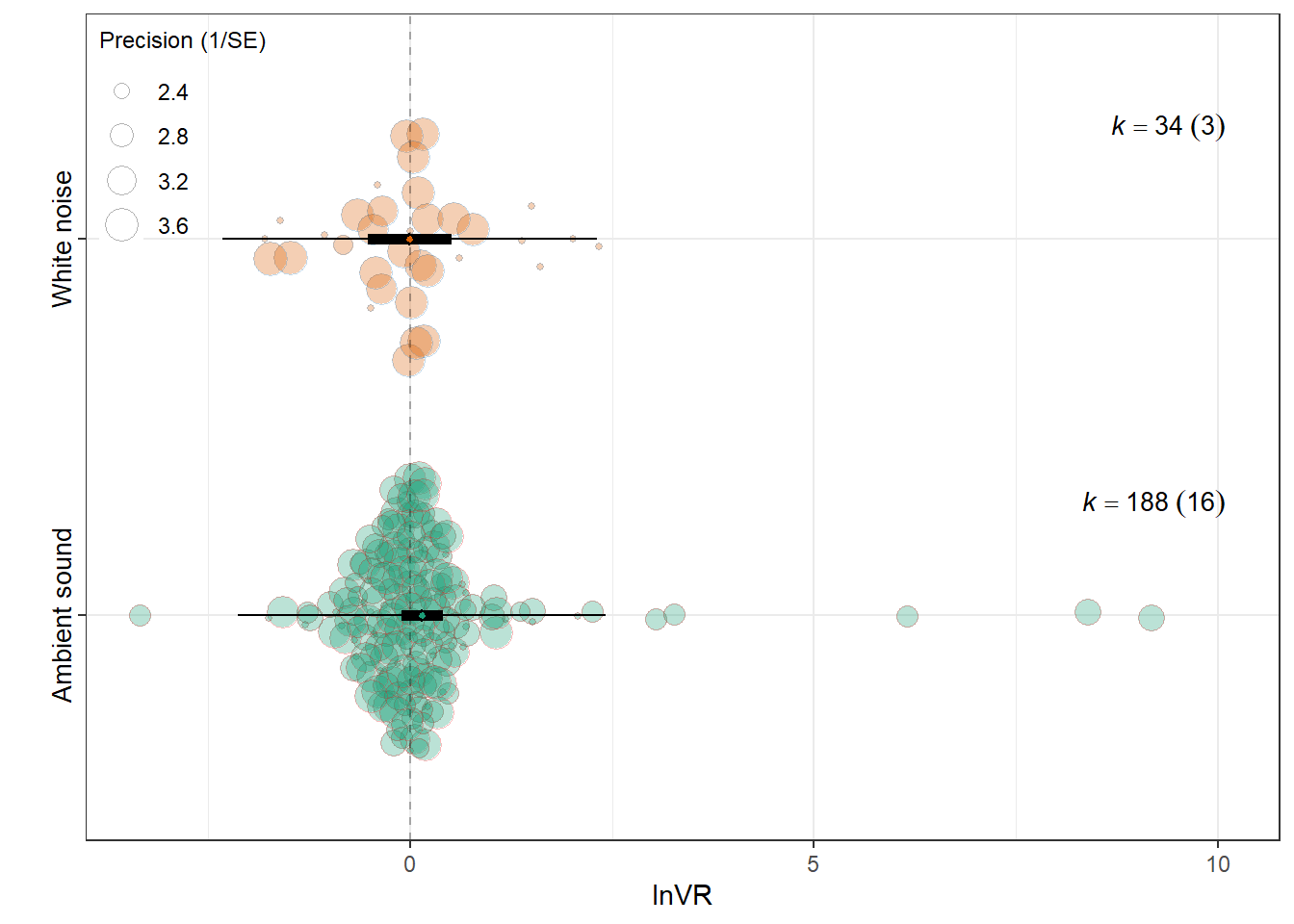
lnCVR
summary(res_cvr$model)
Multivariate Meta-Analysis Model (k = 222; method: REML)
logLik Deviance AIC BIC AICc
-281.7941 563.5883 573.5883 590.5564 573.8687
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0789 0.2808 16 no Study_ID
sigma^2.2 0.5915 0.7691 222 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 220) = 1585.5877, p-val < .0001
Test of Moderators (coefficient 2):
F(df1 = 1, df2 = 220) = 1.7612, p-val = 0.1859
Model Results:
estimate se tval df pval ci.lb
intrcpt 0.0380 0.1192 0.3187 220 0.7503 -0.1969
Control_conditionsWhite noise -0.2540 0.1914 -1.3271 220 0.1859 -0.6313
ci.ub
intrcpt 0.2729
Control_conditionsWhite noise 0.1232
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_cvr$r2 R2_marginal R2_conditional
0.0124 0.1286 res_cvr$plot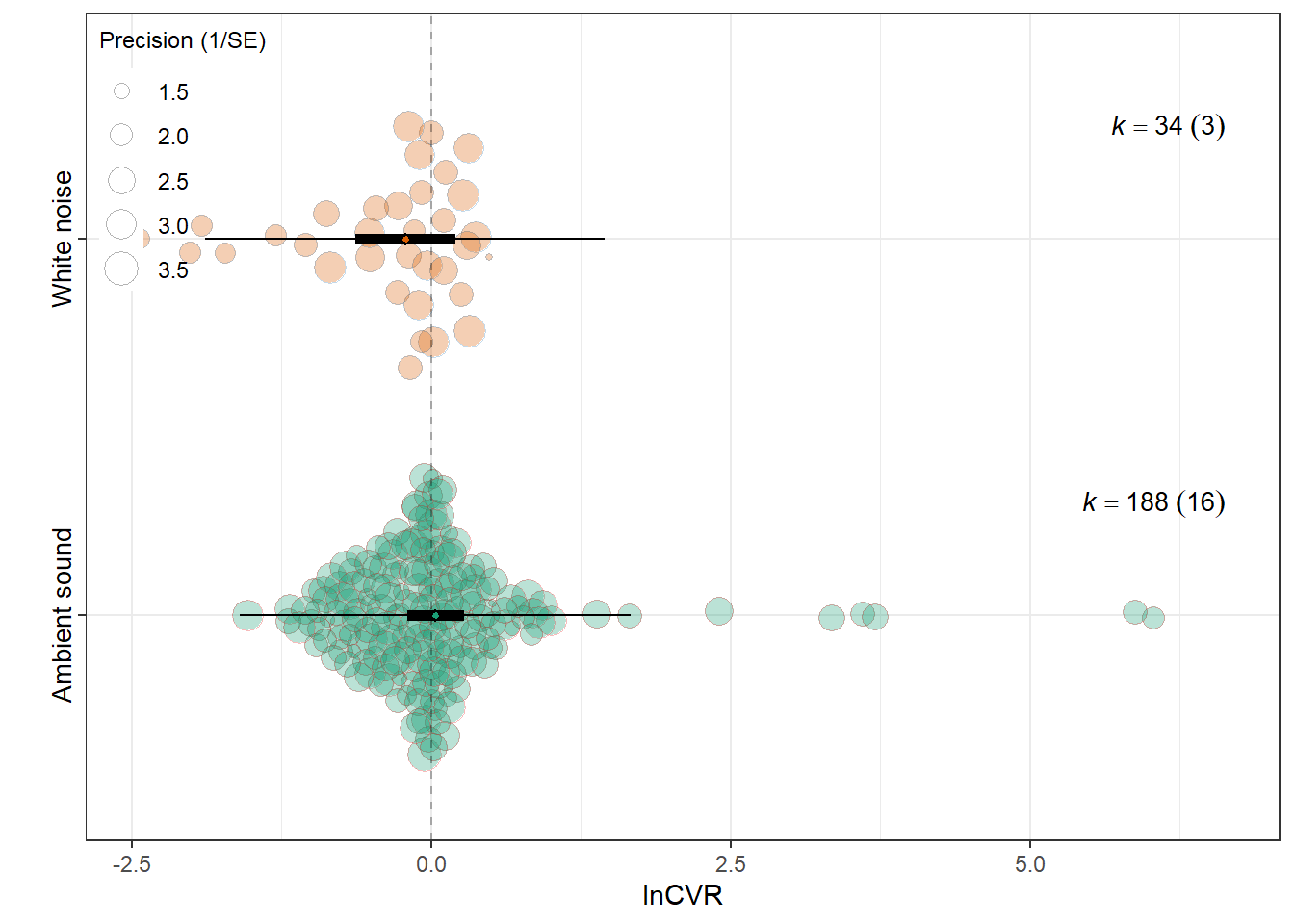
Relative timing
MOD <- "Relative_timing"
MODS <- "~ Relative_timing -1"
# 1) Filter each effect-size dataset (and subset the VCV accordingly)
f_rr <- filter_low_k_single(db_rr, VCV_rr, MOD, min_k = 5)
f_vr <- filter_low_k_single(db_vr, VCV_vr, MOD, min_k = 5)
f_cvr <- filter_low_k_single(db_cvr, VCV_cvr, MOD, min_k = 5)
# 2) Fit the three unimoderator models (same moderator, different yi/vi)
res_rr <- fit_unimod(f_rr$data, f_rr$V, "lnRR", MODS, MOD, expression(lnRR))
res_vr <- fit_unimod(f_vr$data, f_vr$V, "lnVR", MODS, MOD, expression(lnVR))
res_cvr <- fit_unimod(f_cvr$data, f_cvr$V, "lnCVR", MODS, MOD, expression(lnCVR))
# 3) Tukey contrasts (use the filtered data + matched VCV for each dataset)
tuk_rr <- maybe_tukey(dat = f_rr$data, V = f_rr$V, yi = "lnRR", moderator = MOD)
tuk_vr <- maybe_tukey(dat = f_vr$data, V = f_vr$V, yi = "lnVR", moderator = MOD)
tuk_cvr <- maybe_tukey(dat = f_cvr$data, V = f_cvr$V, yi = "lnCVR", moderator = MOD)lnRR
summary(res_rr$model)
Multivariate Meta-Analysis Model (k = 298; method: REML)
logLik Deviance AIC BIC AICc
-238.8518 477.7036 489.7036 511.8254 489.9953
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0495 0.2225 20 no Study_ID
sigma^2.2 0.2274 0.4768 298 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 295) = 5723.3737, p-val < .0001
Test of Moderators (coefficients 1:3):
F(df1 = 3, df2 = 295) = 5.1666, p-val = 0.0017
Model Results:
estimate se tval df pval ci.lb
Relative_timingBefore -0.2276 0.0721 -3.1590 295 0.0017 -0.3695
Relative_timingBoth -0.1965 0.1394 -1.4096 295 0.1597 -0.4707
Relative_timingConcurrent 0.3328 0.2469 1.3475 295 0.1788 -0.1532
ci.ub
Relative_timingBefore -0.0858 **
Relative_timingBoth 0.0778
Relative_timingConcurrent 0.8188
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_rr$r2 R2_marginal R2_conditional
0.0220 0.1969 res_rr$plot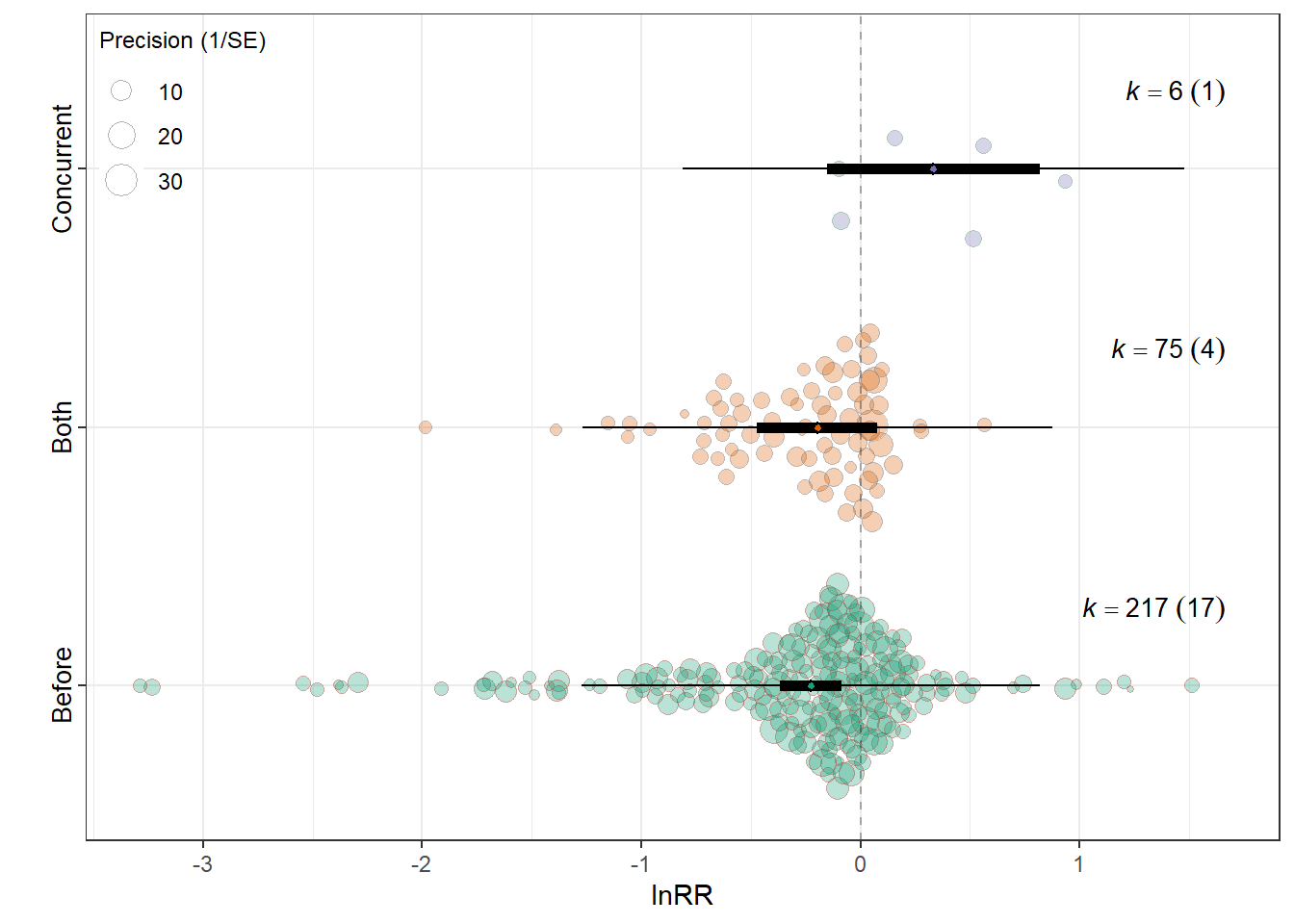
tuk_rr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 0.03119 0.15099 0.207 0.8364
3 - 1 == 0 0.56041 0.24810 2.259 0.0239 *
3 - 2 == 0 0.52922 0.25511 2.074 0.0380 *
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Adjusted p values reported -- none method)lnVR
summary(res_vr$model)
Multivariate Meta-Analysis Model (k = 222; method: REML)
logLik Deviance AIC BIC AICc
-345.0522 690.1044 702.1044 722.4389 702.5006
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0962 0.3102 16 no Study_ID
sigma^2.2 1.2301 1.1091 222 no ES_ID
sigma^2.3 0.0000 0.0001 6 no Strain
Test for Residual Heterogeneity:
QE(df = 219) = 4389.4876, p-val < .0001
Test of Moderators (coefficients 1:3):
F(df1 = 3, df2 = 219) = 0.4882, p-val = 0.6908
Model Results:
estimate se tval df pval ci.lb
Relative_timingBefore 0.1733 0.1445 1.1996 219 0.2316 -0.1114
Relative_timingBoth -0.0209 0.2902 -0.0721 219 0.9426 -0.5928
Relative_timingConcurrent 0.0074 0.5409 0.0136 219 0.9891 -1.0587
ci.ub
Relative_timingBefore 0.4581
Relative_timingBoth 0.5509
Relative_timingConcurrent 1.0735
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_vr$r2 R2_marginal R2_conditional
0.0055 0.0776 res_vr$plot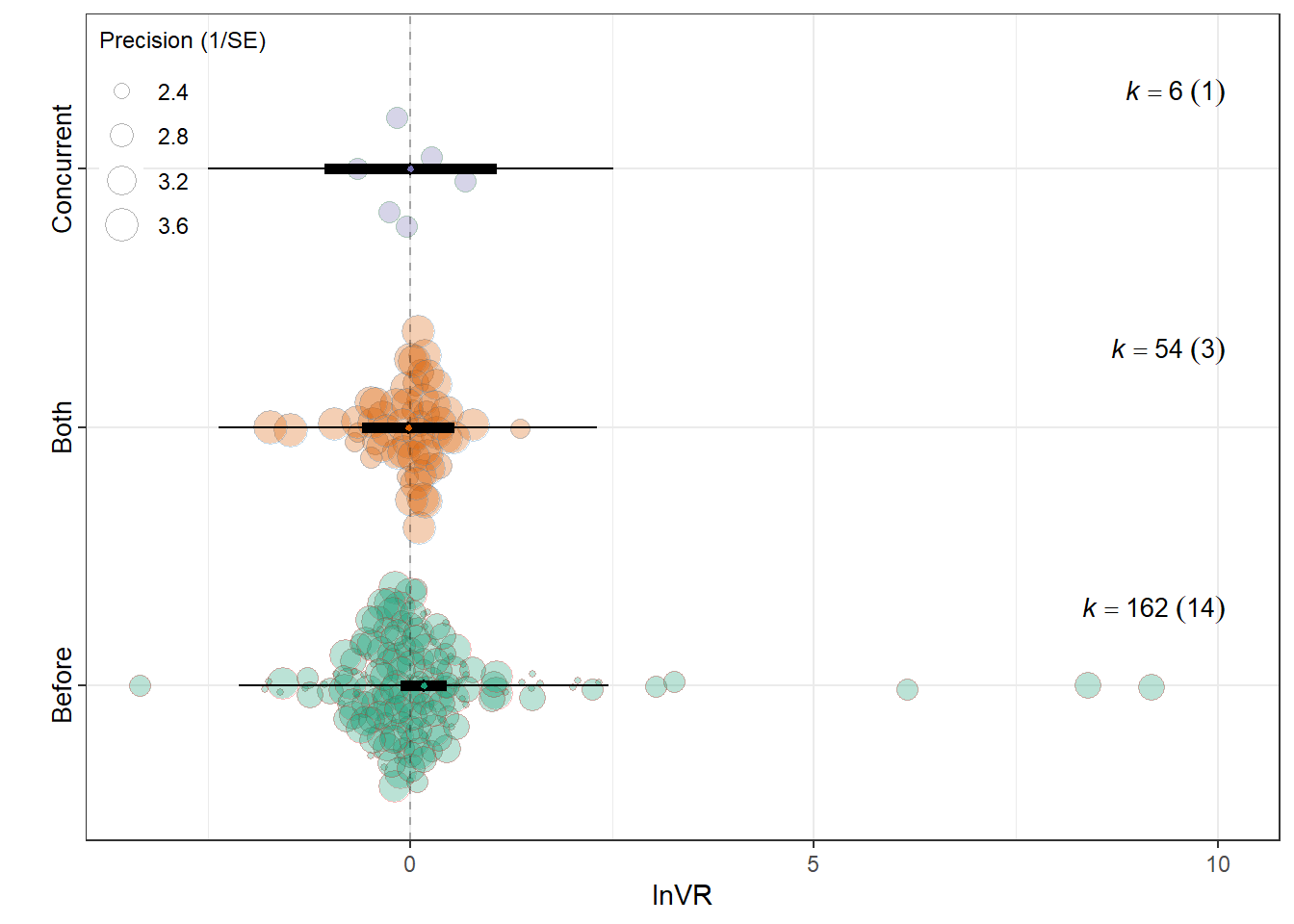
tuk_vr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 -0.19425 0.31647 -0.614 0.539
3 - 1 == 0 -0.16596 0.54716 -0.303 0.762
3 - 2 == 0 0.02829 0.56931 0.050 0.960
(Adjusted p values reported -- none method)lnCVR
summary(res_cvr$model)
Multivariate Meta-Analysis Model (k = 222; method: REML)
logLik Deviance AIC BIC AICc
-281.1298 562.2596 574.2596 594.5940 574.6558
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.1051 0.3241 16 no Study_ID
sigma^2.2 0.5934 0.7703 222 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 219) = 1587.2641, p-val < .0001
Test of Moderators (coefficients 1:3):
F(df1 = 3, df2 = 219) = 0.0805, p-val = 0.9706
Model Results:
estimate se tval df pval ci.lb
Relative_timingBefore 0.0414 0.1351 0.3062 219 0.7598 -0.2248
Relative_timingBoth -0.0876 0.2642 -0.3316 219 0.7405 -0.6082
Relative_timingConcurrent -0.0583 0.4153 -0.1405 219 0.8884 -0.8768
ci.ub
Relative_timingBefore 0.3075
Relative_timingBoth 0.4330
Relative_timingConcurrent 0.7601
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_cvr$r2 R2_marginal R2_conditional
0.0045 0.1543 res_cvr$plot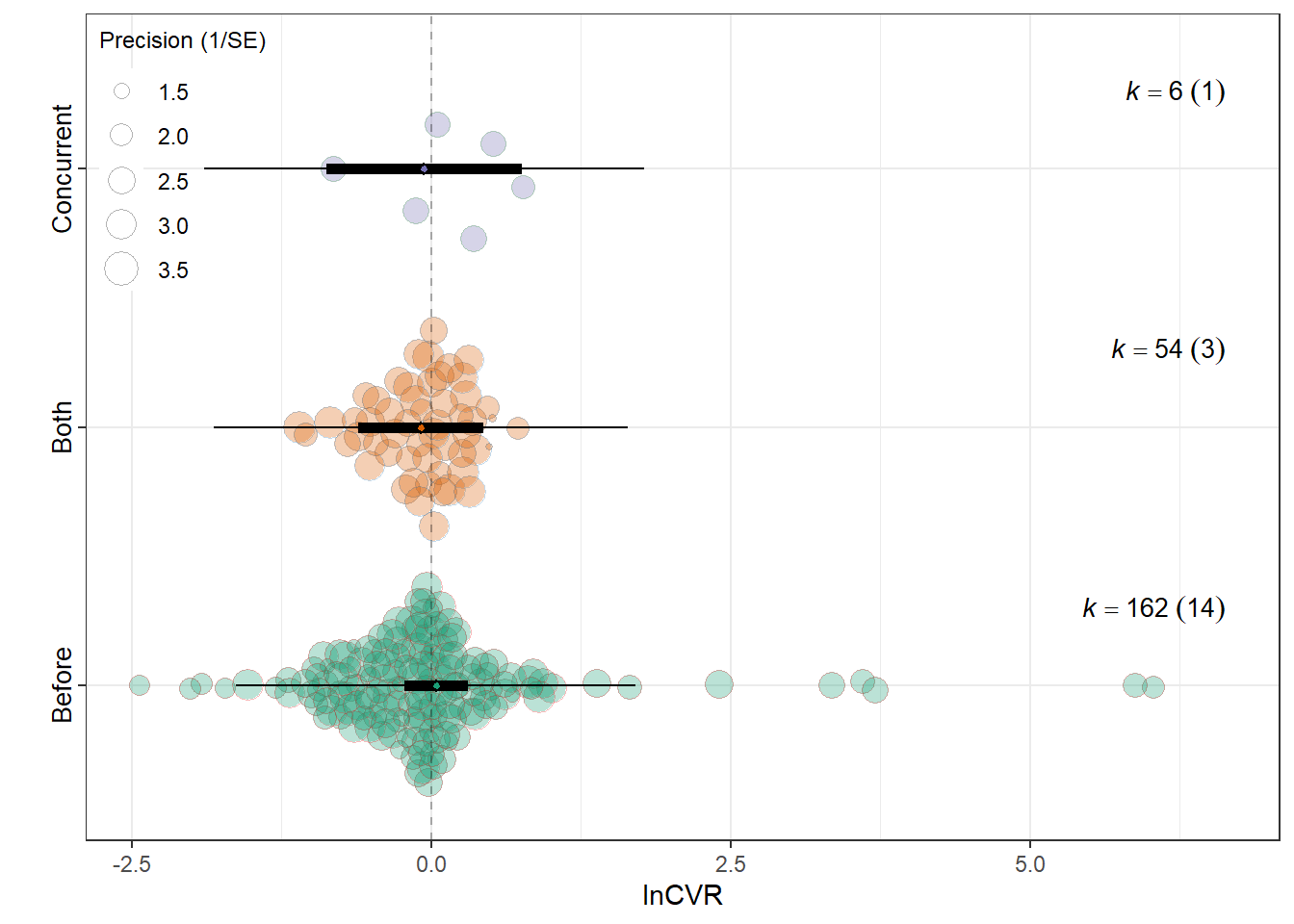
tuk_cvr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 -0.12894 0.28206 -0.457 0.648
3 - 1 == 0 -0.09969 0.41646 -0.239 0.811
3 - 2 == 0 0.02925 0.43058 0.068 0.946
(Adjusted p values reported -- none method)Music genre
MOD <- "Meta_genre"
MODS <- "~ Meta_genre -1"
genre_labels <- c(
"Mixed" = "Mixed",
"Popular Contemporary Music" = "Contemporary",
"Traditional Music / Folk / World" = "World",
"Western Art Music / Orchestral" = "Orchestral"
)
# 1) Filter each effect-size dataset (and subset the VCV accordingly)
f_rr <- filter_low_k_single(db_rr, VCV_rr, MOD, min_k = 5)
f_vr <- filter_low_k_single(db_vr, VCV_vr, MOD, min_k = 5)
f_cvr <- filter_low_k_single(db_cvr, VCV_cvr, MOD, min_k = 5)
# 2) Fit the three unimoderator models (same moderator, different yi/vi)
res_rr <- fit_unimod(f_rr$data, f_rr$V, "lnRR", MODS, MOD, expression(lnRR))
res_vr <- fit_unimod(f_vr$data, f_vr$V, "lnVR", MODS, MOD, expression(lnVR))
res_cvr <- fit_unimod(f_cvr$data, f_cvr$V, "lnCVR", MODS, MOD, expression(lnCVR))
# 2b) Relabel x-axis in the orchard plots (so EPM/OFT/etc. show)
res_rr$plot <- res_rr$plot + scale_x_discrete(labels = genre_labels)
res_vr$plot <- res_vr$plot + scale_x_discrete(labels = genre_labels)
res_cvr$plot <- res_cvr$plot + scale_x_discrete(labels = genre_labels)
# 3) Tukey contrasts (use the filtered data + matched VCV for each dataset)
tuk_rr <- maybe_tukey(dat = f_rr$data, V = f_rr$V, yi = "lnRR", moderator = MOD)
tuk_vr <- maybe_tukey(dat = f_vr$data, V = f_vr$V, yi = "lnVR", moderator = MOD)
tuk_cvr <- maybe_tukey(dat = f_cvr$data, V = f_cvr$V, yi = "lnCVR", moderator = MOD)lnRR
summary(res_rr$model)
Multivariate Meta-Analysis Model (k = 298; method: REML)
logLik Deviance AIC BIC AICc
-239.2750 478.5500 492.5500 518.3351 492.9416
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0497 0.2230 20 no Study_ID
sigma^2.2 0.2304 0.4800 298 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 294) = 5718.2060, p-val < .0001
Test of Moderators (coefficients 1:4):
F(df1 = 4, df2 = 294) = 3.2258, p-val = 0.0130
Model Results:
estimate se tval df
Meta_genreMixed -0.2143 0.1467 -1.4605 294
Meta_genrePopular Contemporary Music -0.1015 0.1917 -0.5297 294
Meta_genreTraditional Music / Folk / World -0.5983 0.2594 -2.3068 294
Meta_genreWestern Art Music / Orchestral -0.1902 0.0766 -2.4812 294
pval ci.lb ci.ub
Meta_genreMixed 0.1452 -0.5030 0.0745
Meta_genrePopular Contemporary Music 0.5968 -0.4787 0.2757
Meta_genreTraditional Music / Folk / World 0.0218 -1.1088 -0.0878 *
Meta_genreWestern Art Music / Orchestral 0.0137 -0.3410 -0.0393 *
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_rr$r2 R2_marginal R2_conditional
0.0232 0.1966 res_rr$plot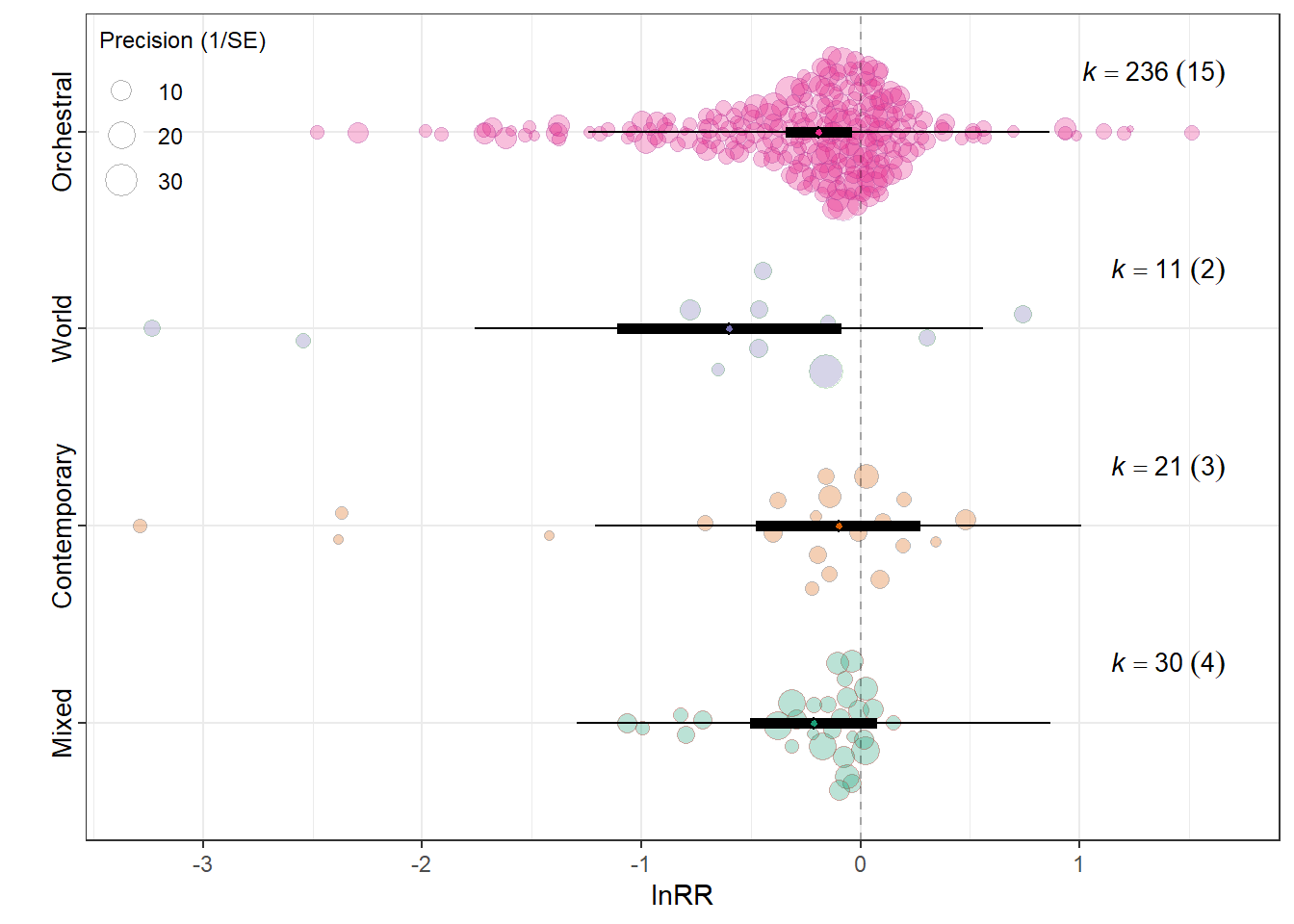
tuk_rr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 0.11277 0.22999 0.490 0.624
3 - 1 == 0 -0.38403 0.29750 -1.291 0.197
4 - 1 == 0 0.02413 0.16005 0.151 0.880
3 - 2 == 0 -0.49680 0.31754 -1.565 0.118
4 - 2 == 0 -0.08864 0.19863 -0.446 0.655
4 - 3 == 0 0.40816 0.26934 1.515 0.130
(Adjusted p values reported -- none method)lnVR
summary(res_vr$model)
Multivariate Meta-Analysis Model (k = 222; method: REML)
logLik Deviance AIC BIC AICc
-338.3565 676.7131 690.7131 714.4045 691.2464
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0000 0.0000 16 no Study_ID
sigma^2.2 1.2200 1.1045 222 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 218) = 4375.2042, p-val < .0001
Test of Moderators (coefficients 1:4):
F(df1 = 4, df2 = 218) = 3.7277, p-val = 0.0059
Model Results:
estimate se tval df
Meta_genreMixed -0.0639 0.2850 -0.2243 218
Meta_genrePopular Contemporary Music 0.9563 0.3203 2.9859 218
Meta_genreTraditional Music / Folk / World 1.1263 0.4627 2.4341 218
Meta_genreWestern Art Music / Orchestral -0.0212 0.1180 -0.1793 218
pval ci.lb ci.ub
Meta_genreMixed 0.8228 -0.6256 0.4978
Meta_genrePopular Contemporary Music 0.0032 0.3251 1.5876 **
Meta_genreTraditional Music / Folk / World 0.0157 0.2143 2.0383 *
Meta_genreWestern Art Music / Orchestral 0.8579 -0.2538 0.2115
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_vr$r2 R2_marginal R2_conditional
0.0841 0.0841 res_vr$plot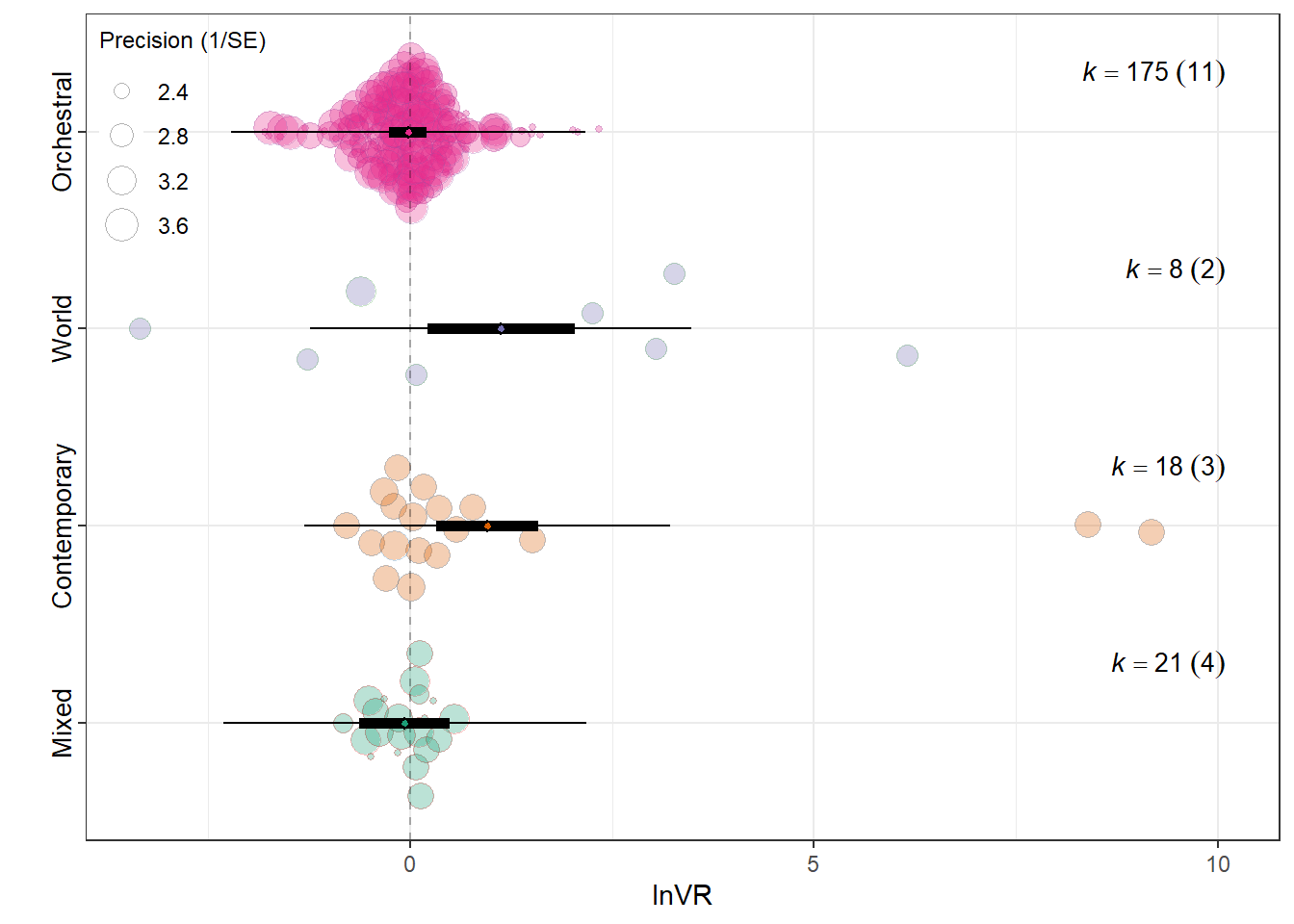
tuk_vr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 1.02023 0.42497 2.401 0.0164 *
3 - 1 == 0 1.19024 0.54342 2.190 0.0285 *
4 - 1 == 0 0.04276 0.30703 0.139 0.8892
3 - 2 == 0 0.17001 0.56173 0.303 0.7622
4 - 2 == 0 -0.97748 0.33961 -2.878 0.0040 **
4 - 3 == 0 -1.14749 0.47737 -2.404 0.0162 *
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Adjusted p values reported -- none method)lnCVR
summary(res_cvr$model)
Multivariate Meta-Analysis Model (k = 222; method: REML)
logLik Deviance AIC BIC AICc
-273.3294 546.6587 560.6587 584.3502 561.1921
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0000 0.0000 16 no Study_ID
sigma^2.2 0.5893 0.7676 222 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 218) = 1563.1637, p-val < .0001
Test of Moderators (coefficients 1:4):
F(df1 = 4, df2 = 218) = 3.9471, p-val = 0.0041
Model Results:
estimate se tval df
Meta_genreMixed -0.0789 0.2334 -0.3381 218
Meta_genrePopular Contemporary Music 0.4492 0.2853 1.5742 218
Meta_genreTraditional Music / Folk / World 1.3149 0.3865 3.4019 218
Meta_genreWestern Art Music / Orchestral -0.1322 0.1071 -1.2342 218
pval ci.lb ci.ub
Meta_genreMixed 0.7356 -0.5390 0.3812
Meta_genrePopular Contemporary Music 0.1169 -0.1132 1.0116
Meta_genreTraditional Music / Folk / World 0.0008 0.5531 2.0767 ***
Meta_genreWestern Art Music / Orchestral 0.2185 -0.3433 0.0789
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_cvr$r2 R2_marginal R2_conditional
0.1359 0.1359 res_cvr$plot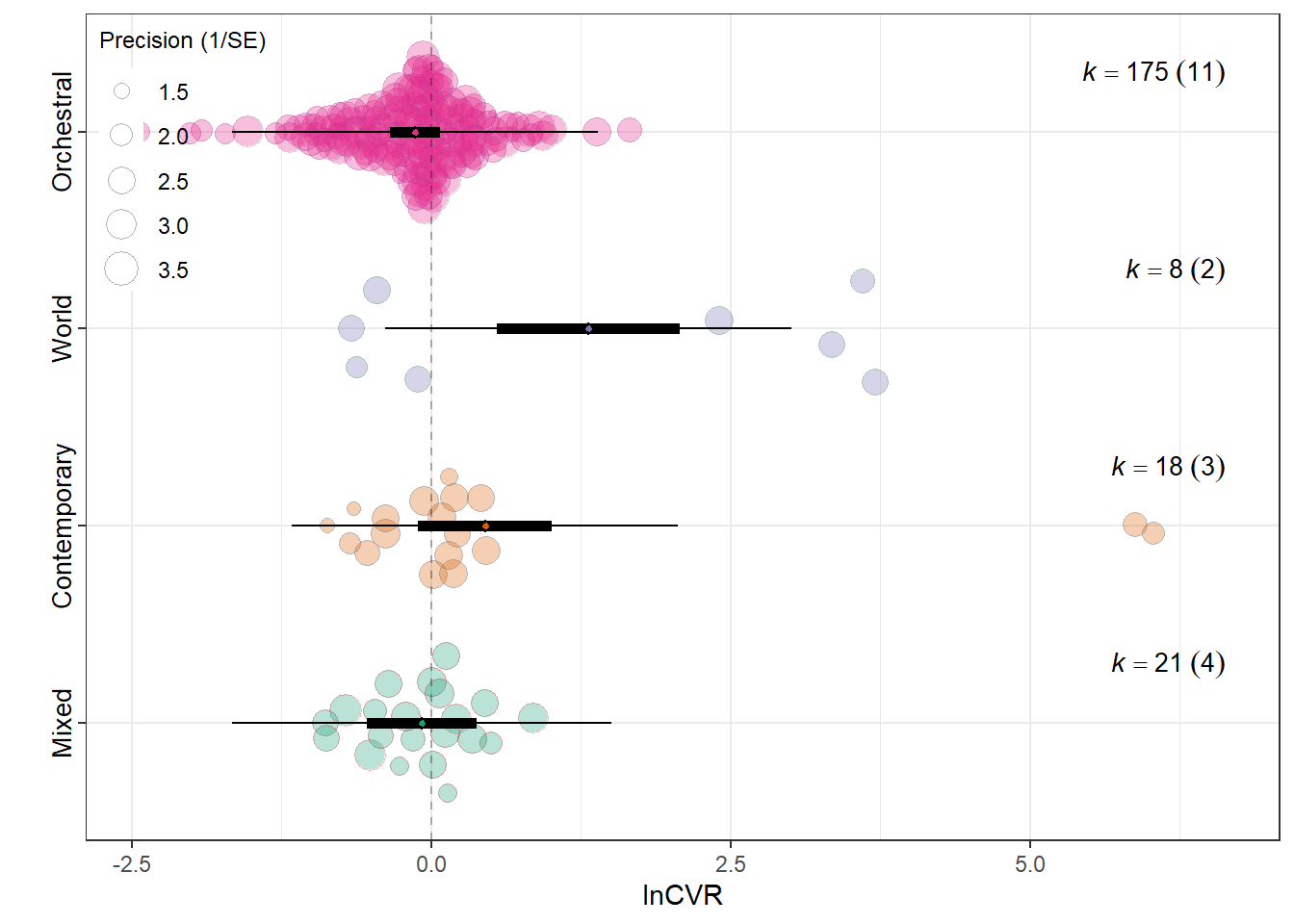
tuk_cvr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 0.52809 0.36032 1.466 0.142754
3 - 1 == 0 1.39383 0.45139 3.088 0.002016 **
4 - 1 == 0 -0.05327 0.25340 -0.210 0.833498
3 - 2 == 0 0.86574 0.47712 1.815 0.069597 .
4 - 2 == 0 -0.58136 0.30018 -1.937 0.052781 .
4 - 3 == 0 -1.44710 0.40050 -3.613 0.000302 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Adjusted p values reported -- none method)Experimental design
MOD <- "Experimental_design"
MODS <- "~ Experimental_design -1"
design_labels<-c(
"Factorial"="Factorial",
"Posttest-Only Control Group"="Posttest",
"Repeated Measures"="Repeated"
)
# 1) Filter each effect-size dataset (and subset the VCV accordingly)
f_rr <- filter_low_k_single(db_rr, VCV_rr, MOD, min_k = 5)
f_vr <- filter_low_k_single(db_vr, VCV_vr, MOD, min_k = 5)
f_cvr <- filter_low_k_single(db_cvr, VCV_cvr, MOD, min_k = 5)
# 2) Fit the three unimoderator models (same moderator, different yi/vi)
res_rr <- fit_unimod(f_rr$data, f_rr$V, "lnRR", MODS, MOD, expression(lnRR))
res_vr <- fit_unimod(f_vr$data, f_vr$V, "lnVR", MODS, MOD, expression(lnVR))
res_cvr <- fit_unimod(f_cvr$data, f_cvr$V, "lnCVR", MODS, MOD, expression(lnCVR))
# 2b) Relabel x-axis in the orchard plots (so EPM/OFT/etc. show)
res_rr$plot <- res_rr$plot + scale_x_discrete(labels = design_labels)
res_vr$plot <- res_vr$plot + scale_x_discrete(labels = design_labels)
res_cvr$plot <- res_cvr$plot + scale_x_discrete(labels = design_labels)
# 3) Tukey contrasts (use the filtered data + matched VCV for each dataset)
tuk_rr <- maybe_tukey(dat = f_rr$data, V = f_rr$V, yi = "lnRR", moderator = MOD)
tuk_vr <- maybe_tukey(dat = f_vr$data, V = f_vr$V, yi = "lnVR", moderator = MOD)
tuk_cvr <- maybe_tukey(dat = f_cvr$data, V = f_cvr$V, yi = "lnCVR", moderator = MOD)lnRR
summary(res_rr$model)
Multivariate Meta-Analysis Model (k = 298; method: REML)
logLik Deviance AIC BIC AICc
-239.2296 478.4593 490.4593 512.5811 490.7509
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0442 0.2102 20 no Study_ID
sigma^2.2 0.2297 0.4793 298 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 295) = 5684.0603, p-val < .0001
Test of Moderators (coefficients 1:3):
F(df1 = 3, df2 = 295) = 5.0020, p-val = 0.0021
Model Results:
estimate se tval df
Experimental_designFactorial -0.1772 0.0704 -2.5178 295
Experimental_designPosttest-Only Control Group -0.3808 0.1066 -3.5735 295
Experimental_designRepeated Measures -0.1624 0.2051 -0.7915 295
pval ci.lb ci.ub
Experimental_designFactorial 0.0123 -0.3157 -0.0387 *
Experimental_designPosttest-Only Control Group 0.0004 -0.5905 -0.1711 ***
Experimental_designRepeated Measures 0.4293 -0.5660 0.2413
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_rr$r2 R2_marginal R2_conditional
0.0224 0.1801 res_rr$plot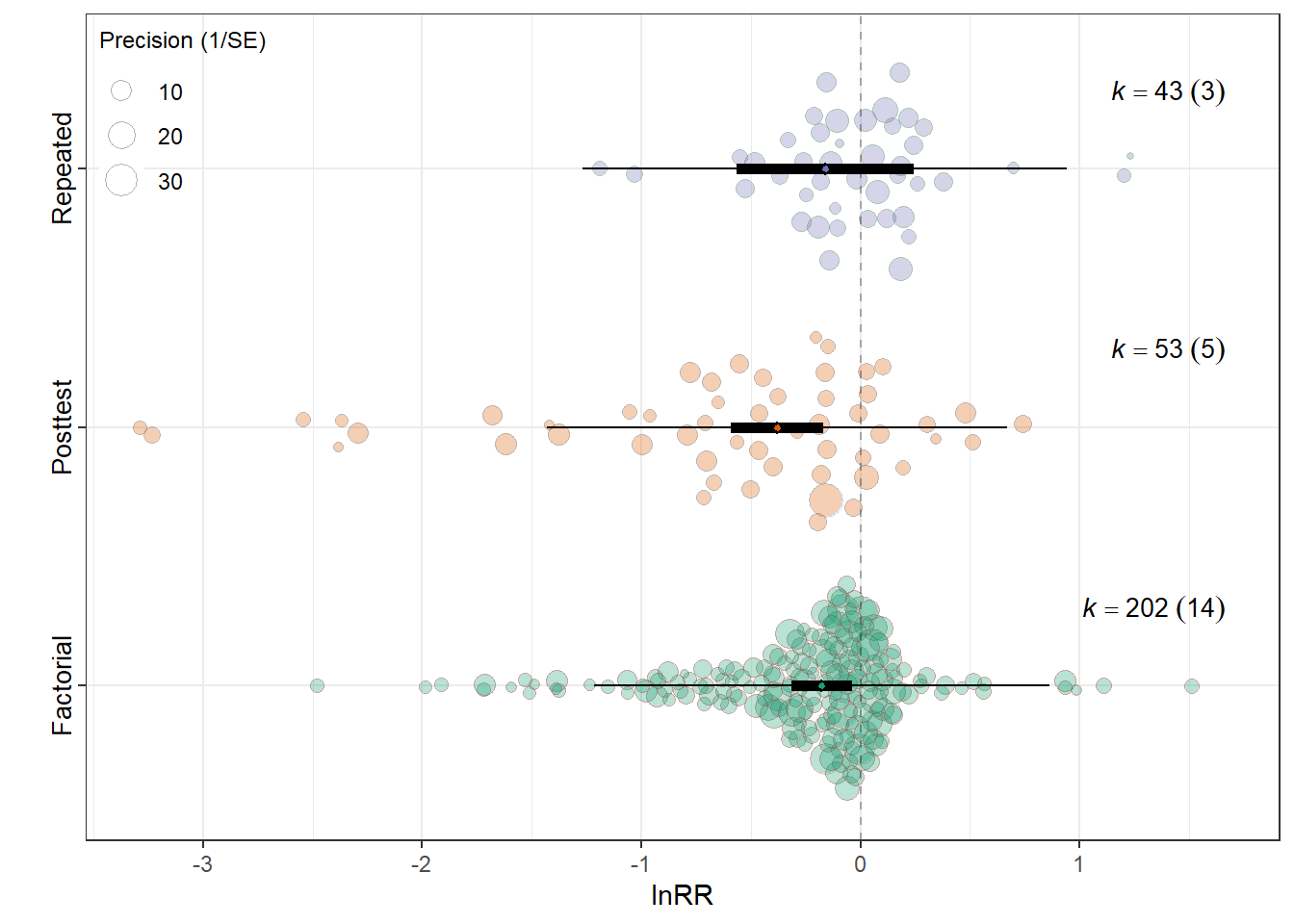
tuk_rr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 -0.20356 0.10332 -1.970 0.0488 *
3 - 1 == 0 0.01485 0.21686 0.068 0.9454
3 - 2 == 0 0.21841 0.23115 0.945 0.3447
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Adjusted p values reported -- none method)lnVR
summary(res_vr$model)
Multivariate Meta-Analysis Model (k = 222; method: REML)
logLik Deviance AIC BIC AICc
-341.4846 682.9693 694.9693 715.3037 695.3655
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0000 0.0001 16 no Study_ID
sigma^2.2 1.2318 1.1099 222 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 219) = 4376.1207, p-val < .0001
Test of Moderators (coefficients 1:3):
F(df1 = 3, df2 = 219) = 3.6274, p-val = 0.0138
Model Results:
estimate se tval df
Experimental_designFactorial 0.0225 0.1185 0.1899 219
Experimental_designPosttest-Only Control Group 0.6459 0.2007 3.2188 219
Experimental_designRepeated Measures -0.2031 0.3078 -0.6598 219
pval ci.lb ci.ub
Experimental_designFactorial 0.8496 -0.2110 0.2560
Experimental_designPosttest-Only Control Group 0.0015 0.2504 1.0414 **
Experimental_designRepeated Measures 0.5101 -0.8097 0.4035
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_vr$r2 R2_marginal R2_conditional
0.056 0.056 res_vr$plot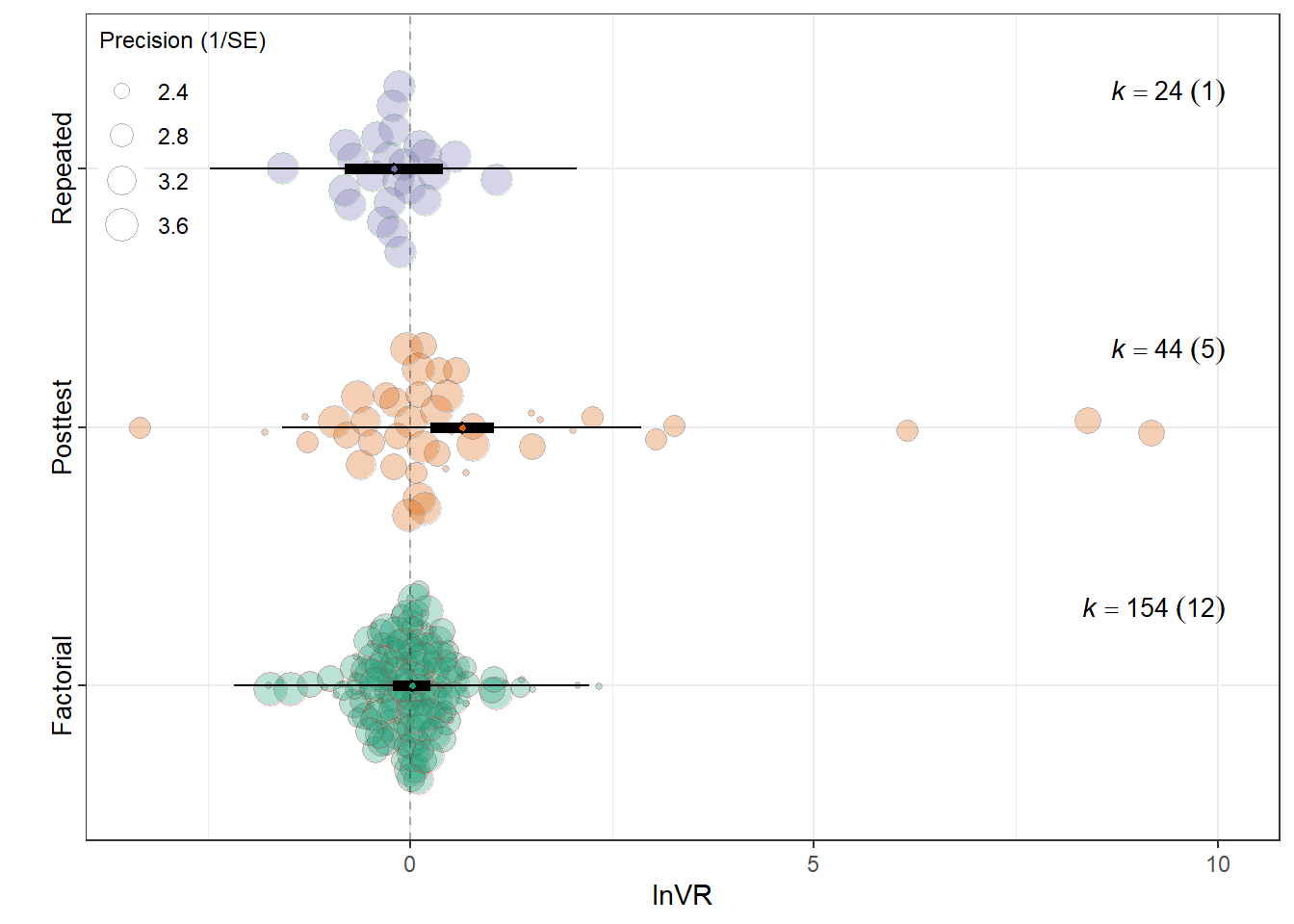
tuk_vr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 0.6234 0.2173 2.869 0.00412 **
3 - 1 == 0 -0.2256 0.3298 -0.684 0.49399
3 - 2 == 0 -0.8490 0.3674 -2.311 0.02085 *
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Adjusted p values reported -- none method)lnCVR
summary(res_cvr$model)
Multivariate Meta-Analysis Model (k = 222; method: REML)
logLik Deviance AIC BIC AICc
-280.1420 560.2840 572.2840 592.6184 572.6802
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0560 0.2366 16 no Study_ID
sigma^2.2 0.5990 0.7739 222 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 219) = 1585.4977, p-val < .0001
Test of Moderators (coefficients 1:3):
F(df1 = 3, df2 = 219) = 0.7164, p-val = 0.5431
Model Results:
estimate se tval df
Experimental_designFactorial -0.0239 0.1243 -0.1926 219
Experimental_designPosttest-Only Control Group 0.2250 0.1831 1.2286 219
Experimental_designRepeated Measures -0.1655 0.3664 -0.4516 219
pval ci.lb ci.ub
Experimental_designFactorial 0.8475 -0.2689 0.2210
Experimental_designPosttest-Only Control Group 0.2205 -0.1359 0.5859
Experimental_designRepeated Measures 0.6520 -0.8876 0.5567
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_cvr$r2 R2_marginal R2_conditional
0.0200 0.1038 res_cvr$plot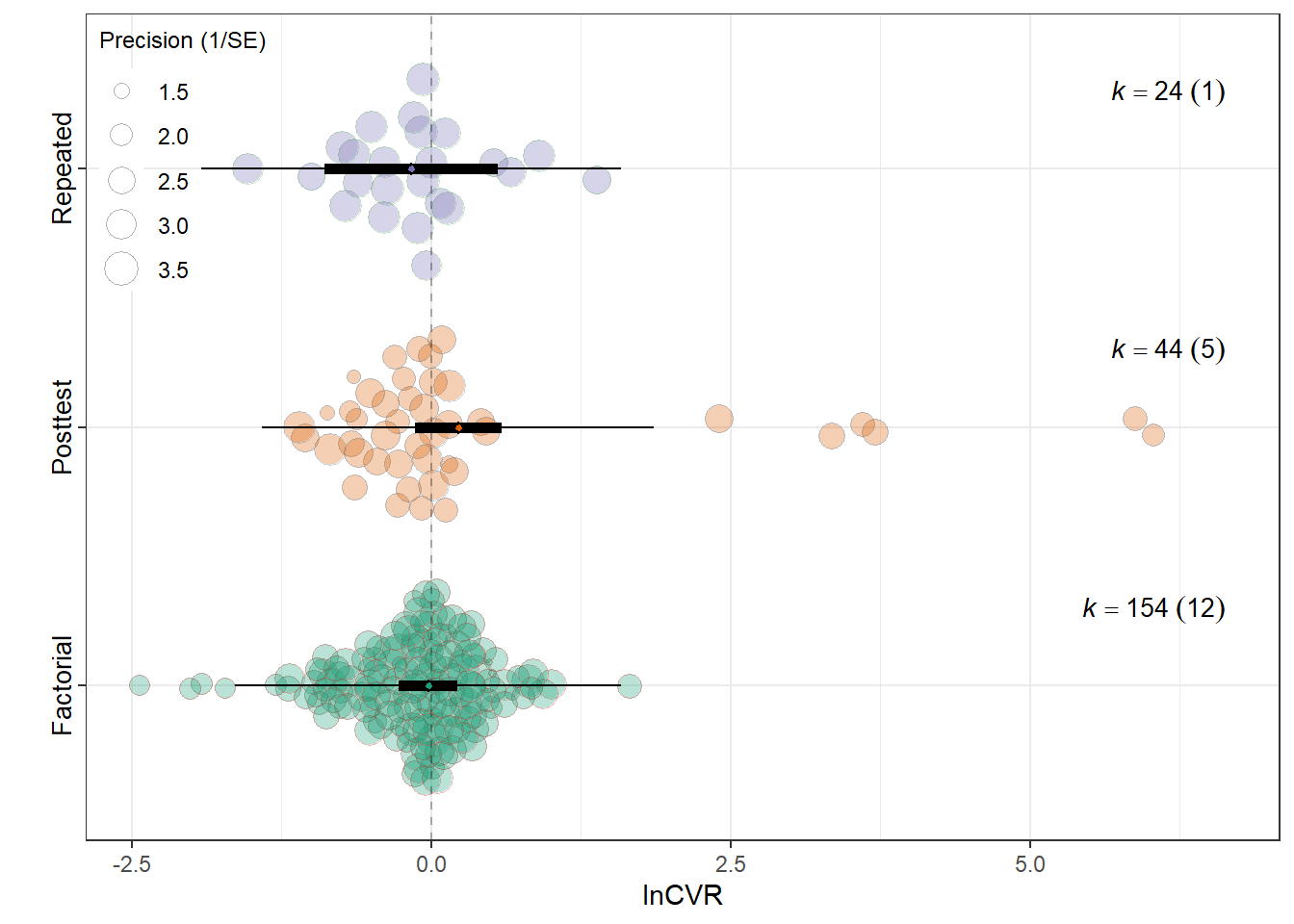
tuk_cvr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 0.2489 0.1821 1.367 0.172
3 - 1 == 0 -0.1415 0.3869 -0.366 0.715
3 - 2 == 0 -0.3904 0.4096 -0.953 0.341
(Adjusted p values reported -- none method)Sex
MOD <- "Sex"
MODS <- "~ Sex"
f_rr <- filter_low_k_single(db_rr, VCV_rr, MOD, min_k = 5)
f_vr <- filter_low_k_single(db_vr, VCV_vr, MOD, min_k = 5)
f_cvr <- filter_low_k_single(db_cvr, VCV_cvr, MOD, min_k = 5)
res_rr <- fit_unimod(f_rr$data, f_rr$V, "lnRR", MODS, MOD, expression(lnRR))
res_vr <- fit_unimod(f_vr$data, f_vr$V, "lnVR", MODS, MOD, expression(lnVR))
res_cvr <- fit_unimod(f_cvr$data, f_cvr$V, "lnCVR", MODS, MOD, expression(lnCVR))lnRR
summary(res_rr$model)
Multivariate Meta-Analysis Model (k = 295; method: REML)
logLik Deviance AIC BIC AICc
-238.9455 477.8911 487.8911 506.2920 488.1002
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0420 0.2050 19 no Study_ID
sigma^2.2 0.2300 0.4796 295 no ES_ID
sigma^2.3 0.0096 0.0981 6 no Strain
Test for Residual Heterogeneity:
QE(df = 293) = 5730.3718, p-val < .0001
Test of Moderators (coefficient 2):
F(df1 = 1, df2 = 293) = 2.7371, p-val = 0.0991
Model Results:
estimate se tval df pval ci.lb ci.ub
intrcpt -0.3100 0.1015 -3.0543 293 0.0025 -0.5098 -0.1103 **
SexMale 0.1529 0.0924 1.6544 293 0.0991 -0.0290 0.3348 .
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_rr$r2 R2_marginal R2_conditional
0.0194 0.1993 res_rr$plot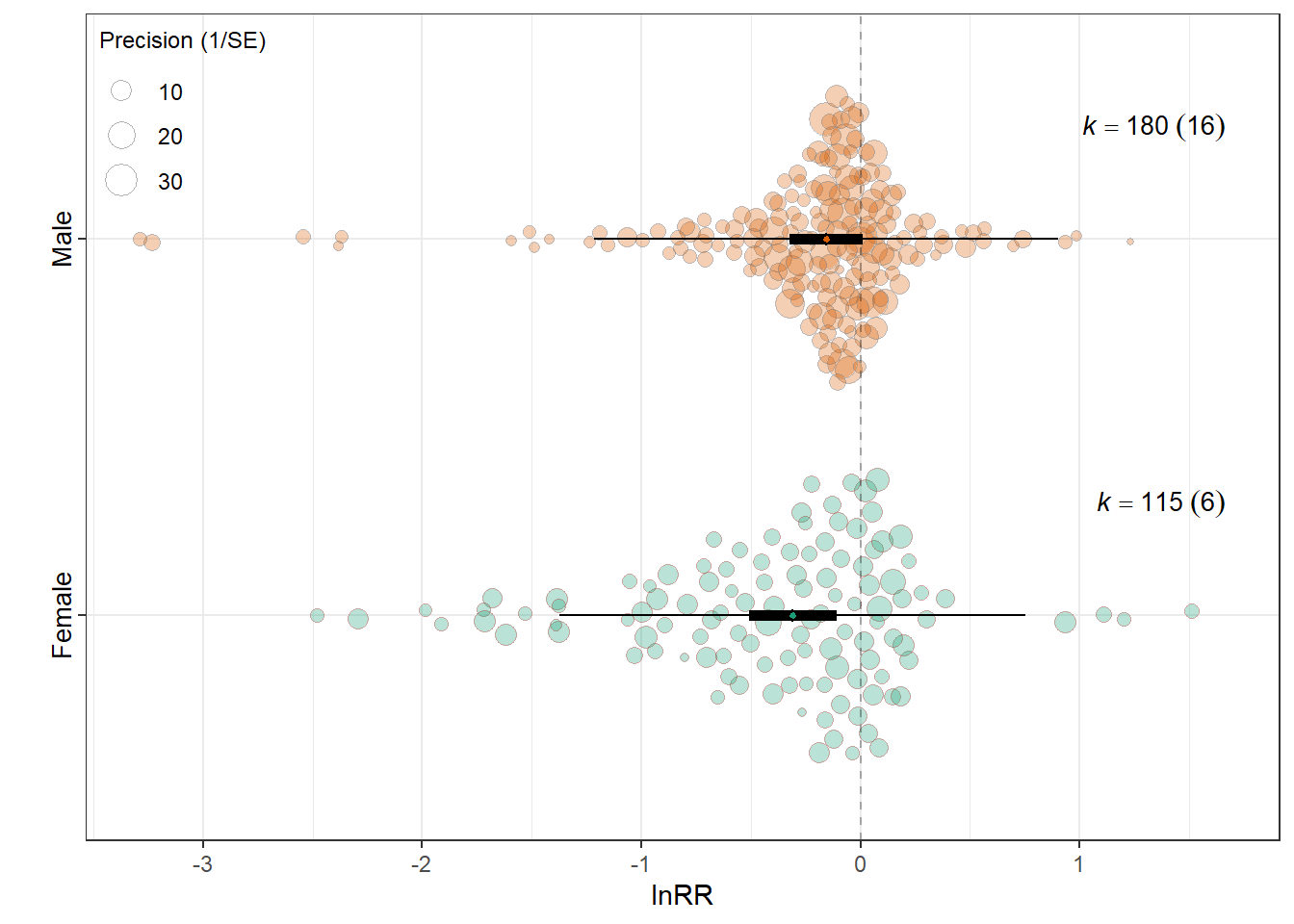
lnVR
summary(res_vr$model)
Multivariate Meta-Analysis Model (k = 219; method: REML)
logLik Deviance AIC BIC AICc
-342.8395 685.6790 695.6790 712.5785 695.9634
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0900 0.3001 15 no Study_ID
sigma^2.2 1.2406 1.1138 219 no ES_ID
sigma^2.3 0.0000 0.0001 6 no Strain
Test for Residual Heterogeneity:
QE(df = 217) = 4383.5230, p-val < .0001
Test of Moderators (coefficient 2):
F(df1 = 1, df2 = 217) = 0.3114, p-val = 0.5774
Model Results:
estimate se tval df pval ci.lb ci.ub
intrcpt 0.0666 0.2034 0.3275 217 0.7436 -0.3342 0.4675
SexMale 0.1276 0.2287 0.5580 217 0.5774 -0.3232 0.5784
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_vr$r2 R2_marginal R2_conditional
0.0030 0.0704 res_vr$plot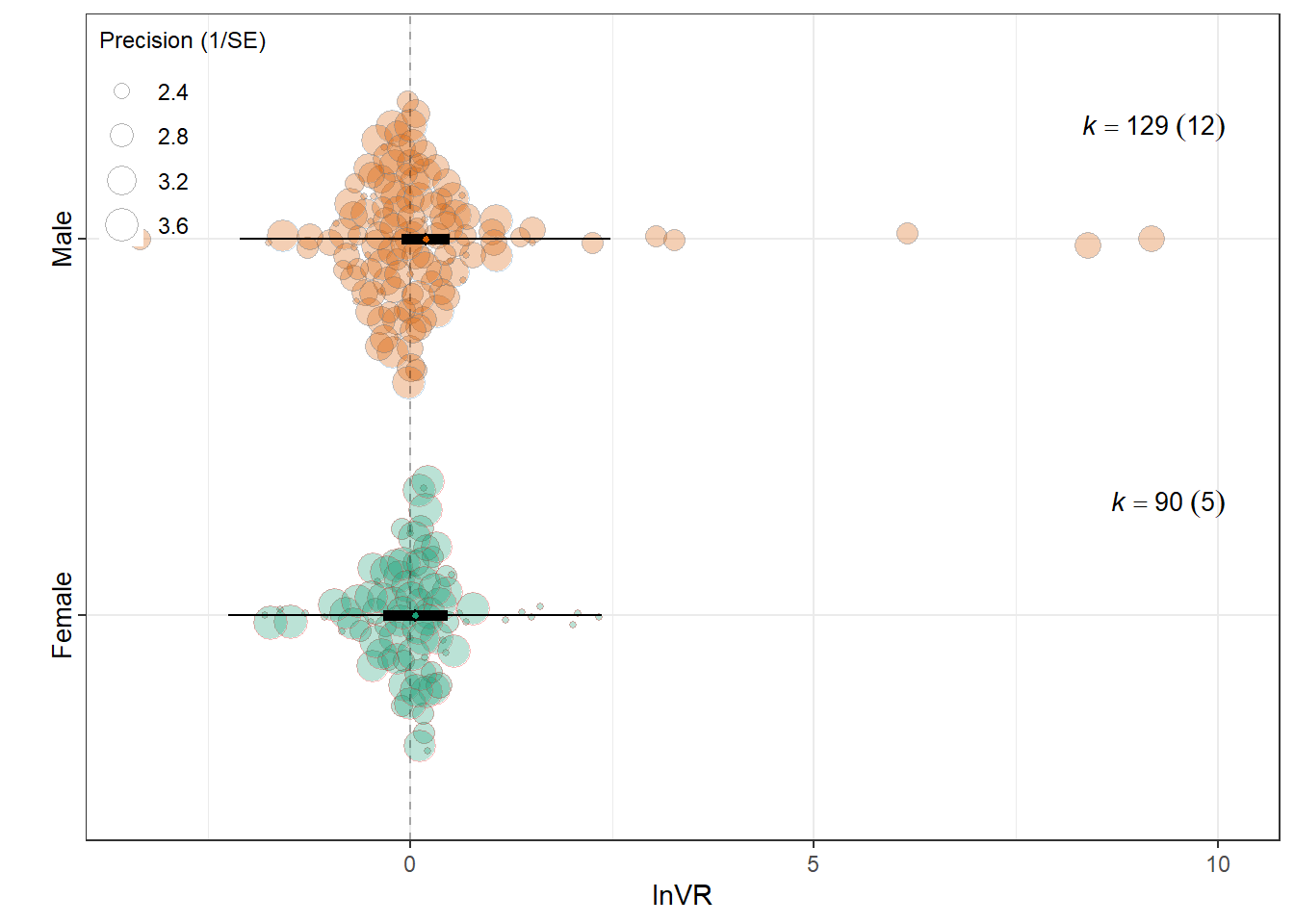
lnCVR
summary(res_cvr$model)
Multivariate Meta-Analysis Model (k = 219; method: REML)
logLik Deviance AIC BIC AICc
-278.1954 556.3907 566.3907 583.2902 566.6751
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0692 0.2630 15 no Study_ID
sigma^2.2 0.5972 0.7728 219 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 217) = 1572.7582, p-val < .0001
Test of Moderators (coefficient 2):
F(df1 = 1, df2 = 217) = 2.0491, p-val = 0.1537
Model Results:
estimate se tval df pval ci.lb ci.ub
intrcpt -0.1386 0.1715 -0.8083 217 0.4198 -0.4766 0.1994
SexMale 0.2616 0.1827 1.4315 217 0.1537 -0.0986 0.6217
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_cvr$r2 R2_marginal R2_conditional
0.0244 0.1256 res_cvr$plot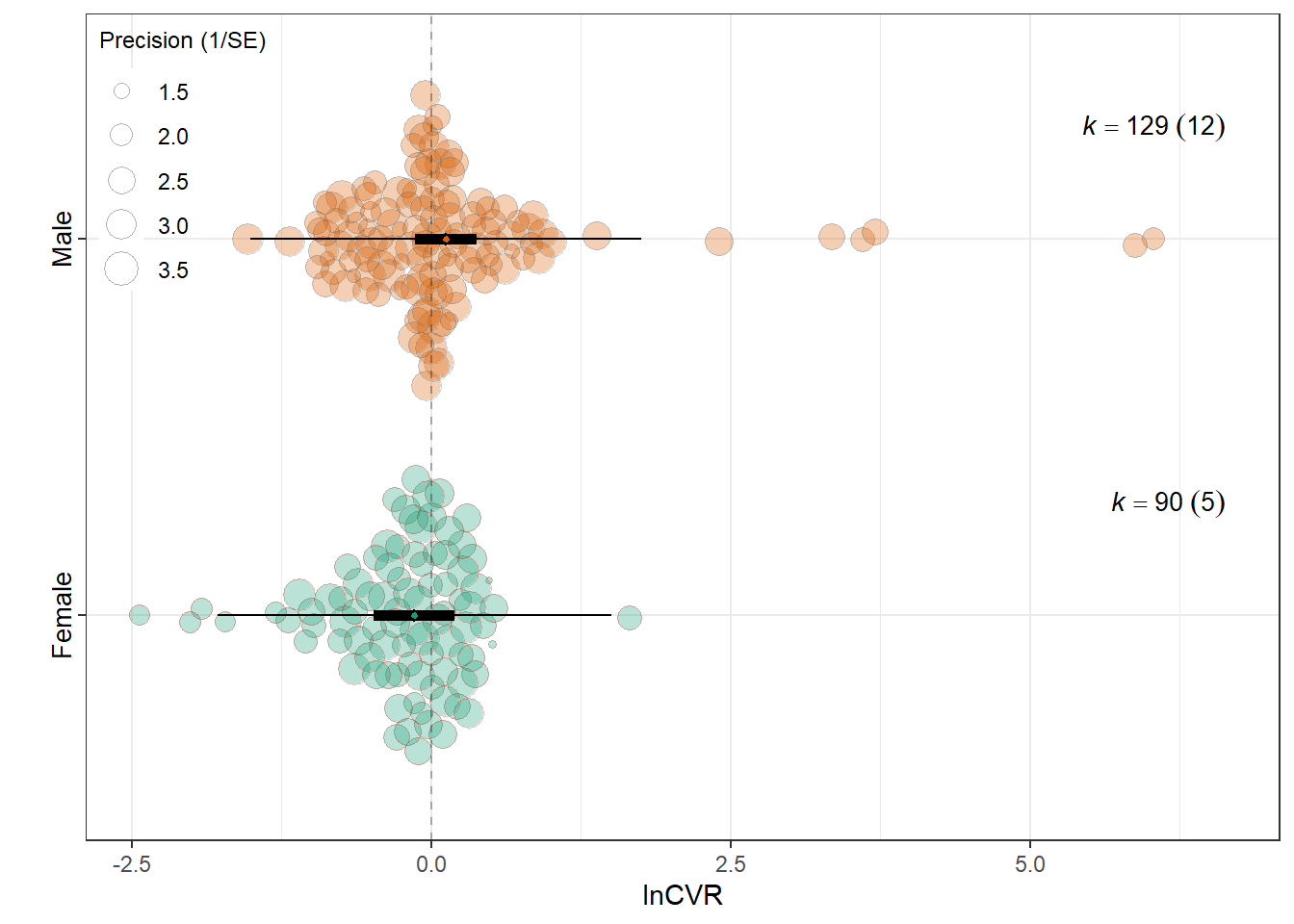
Exposure duration
Music_exposure_duration
MOD <- "Music_exposure_duration"
MODS <- "~ Music_exposure_duration -1"
# 1) Filter each effect-size dataset (and subset the VCV accordingly)
f_rr <- filter_low_k_single(db_rr, VCV_rr, MOD, min_k = 5)
f_vr <- filter_low_k_single(db_vr, VCV_vr, MOD, min_k = 5)
f_cvr <- filter_low_k_single(db_cvr, VCV_cvr, MOD, min_k = 5)
# 2) Fit the three unimoderator models (same moderator, different yi/vi)
res_rr <- fit_unimod(f_rr$data, f_rr$V, "lnRR", MODS, MOD, expression(lnRR))
res_vr <- fit_unimod(f_vr$data, f_vr$V, "lnVR", MODS, MOD, expression(lnVR))
res_cvr <- fit_unimod(f_cvr$data, f_cvr$V, "lnCVR", MODS, MOD, expression(lnCVR))
# 3) Tukey contrasts (use the filtered data + matched VCV for each dataset)
tuk_rr <- maybe_tukey(dat = f_rr$data, V = f_rr$V, yi = "lnRR", moderator = MOD)
tuk_vr <- maybe_tukey(dat = f_vr$data, V = f_vr$V, yi = "lnVR", moderator = MOD)
tuk_cvr <- maybe_tukey(dat = f_cvr$data, V = f_cvr$V, yi = "lnCVR", moderator = MOD)lnRR
summary(res_rr$model)
Multivariate Meta-Analysis Model (k = 297; method: REML)
logLik Deviance AIC BIC AICc
-239.5997 479.1994 491.1994 513.3009 491.4921
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0512 0.2263 20 no Study_ID
sigma^2.2 0.2319 0.4816 297 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 294) = 5777.2420, p-val < .0001
Test of Moderators (coefficients 1:3):
F(df1 = 3, df2 = 294) = 3.8341, p-val = 0.0102
Model Results:
estimate se tval df pval ci.lb
Music_exposure_durationAcute -0.2660 0.1118 -2.3798 294 0.0180 -0.4860
Music_exposure_durationMedium -0.0642 0.1396 -0.4599 294 0.6459 -0.3389
Music_exposure_durationShort -0.2495 0.1052 -2.3722 294 0.0183 -0.4566
ci.ub
Music_exposure_durationAcute -0.0460 *
Music_exposure_durationMedium 0.2105
Music_exposure_durationShort -0.0425 *
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_rr$r2 R2_marginal R2_conditional
0.0216 0.1986 res_rr$plot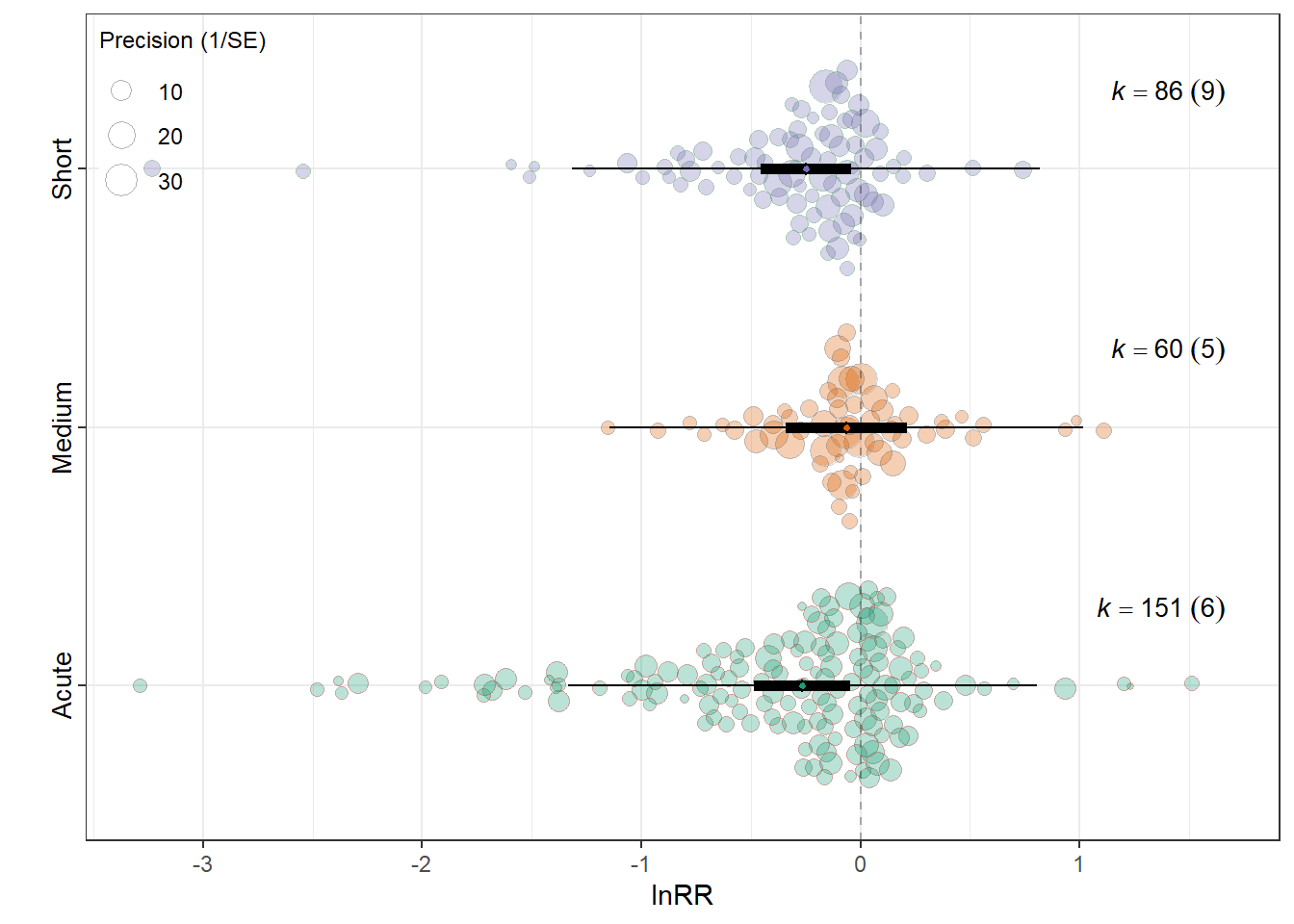
tuk_rr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 0.20184 0.17884 1.129 0.259
3 - 1 == 0 0.01649 0.15350 0.107 0.914
3 - 2 == 0 -0.18535 0.17479 -1.060 0.289
(Adjusted p values reported -- none method)lnVR
summary(res_vr$model)
Multivariate Meta-Analysis Model (k = 222; method: REML)
logLik Deviance AIC BIC AICc
-344.5642 689.1285 701.1285 721.4629 701.5247
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.1181 0.3437 16 no Study_ID
sigma^2.2 1.2224 1.1056 222 no ES_ID
sigma^2.3 0.0000 0.0001 6 no Strain
Test for Residual Heterogeneity:
QE(df = 219) = 4390.2022, p-val < .0001
Test of Moderators (coefficients 1:3):
F(df1 = 3, df2 = 219) = 0.5542, p-val = 0.6458
Model Results:
estimate se tval df pval ci.lb
Music_exposure_durationAcute 0.2718 0.2202 1.2345 219 0.2183 -0.1621
Music_exposure_durationMedium 0.0052 0.2628 0.0198 219 0.9843 -0.5127
Music_exposure_durationShort 0.0873 0.2348 0.3717 219 0.7105 -0.3754
ci.ub
Music_exposure_durationAcute 0.7057
Music_exposure_durationMedium 0.5231
Music_exposure_durationShort 0.5499
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_vr$r2 R2_marginal R2_conditional
0.0101 0.0973 res_vr$plot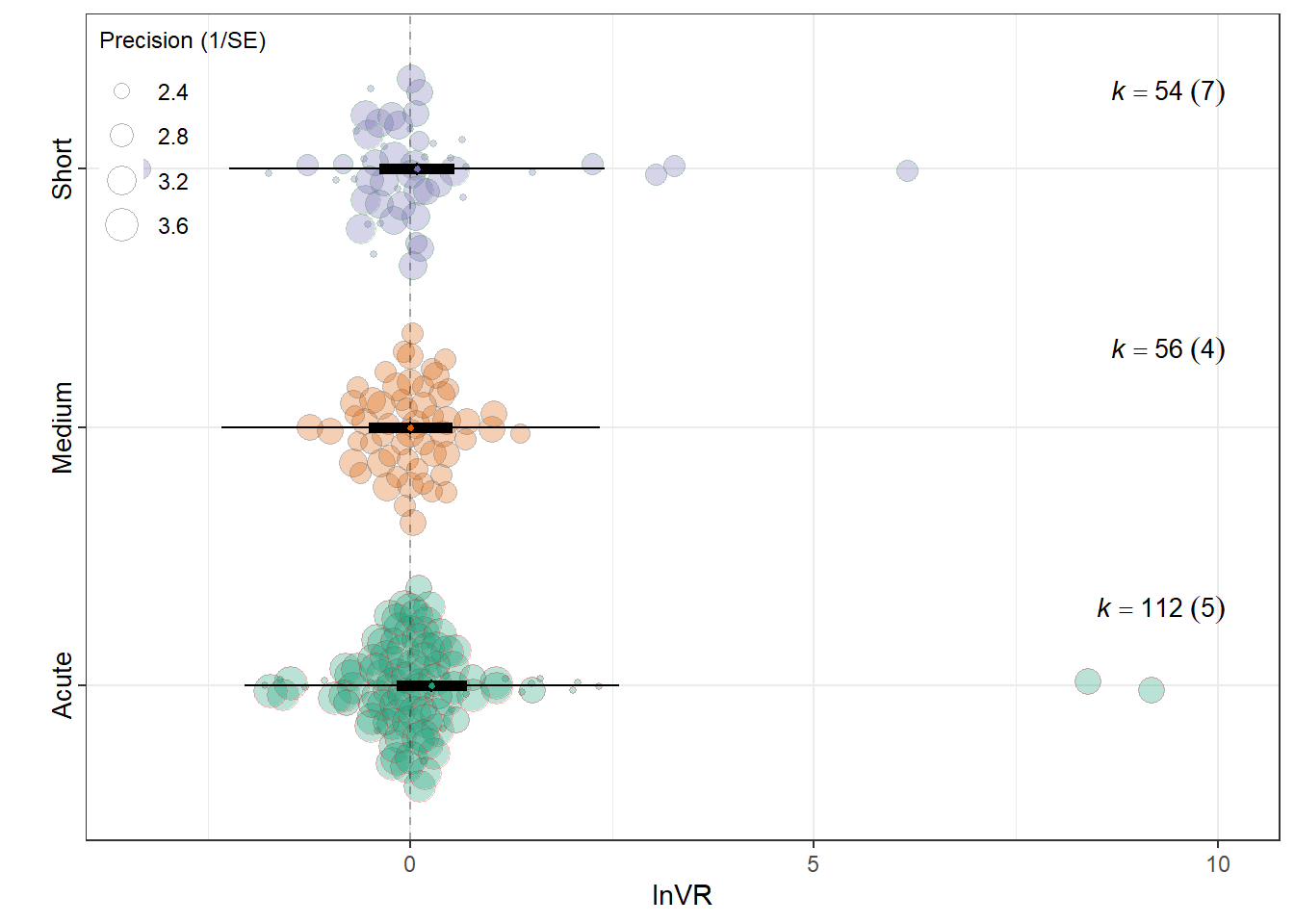
tuk_vr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 -0.26659 0.34281 -0.778 0.437
3 - 1 == 0 -0.18452 0.32184 -0.573 0.566
3 - 2 == 0 0.08207 0.35236 0.233 0.816
(Adjusted p values reported -- none method)lnCVR
summary(res_cvr$model)
Multivariate Meta-Analysis Model (k = 222; method: REML)
logLik Deviance AIC BIC AICc
-280.5247 561.0493 573.0493 593.3838 573.4456
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.1365 0.3695 16 no Study_ID
sigma^2.2 0.5889 0.7674 222 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 219) = 1588.0953, p-val < .0001
Test of Moderators (coefficients 1:3):
F(df1 = 3, df2 = 219) = 0.0367, p-val = 0.9906
Model Results:
estimate se tval df pval ci.lb
Music_exposure_durationAcute 0.0200 0.2204 0.0907 219 0.9278 -0.4143
Music_exposure_durationMedium -0.0405 0.2568 -0.1579 219 0.8747 -0.5466
Music_exposure_durationShort 0.0610 0.2200 0.2771 219 0.7820 -0.3726
ci.ub
Music_exposure_durationAcute 0.4543
Music_exposure_durationMedium 0.4655
Music_exposure_durationShort 0.4945
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_cvr$r2 R2_marginal R2_conditional
0.0018 0.1897 res_cvr$plot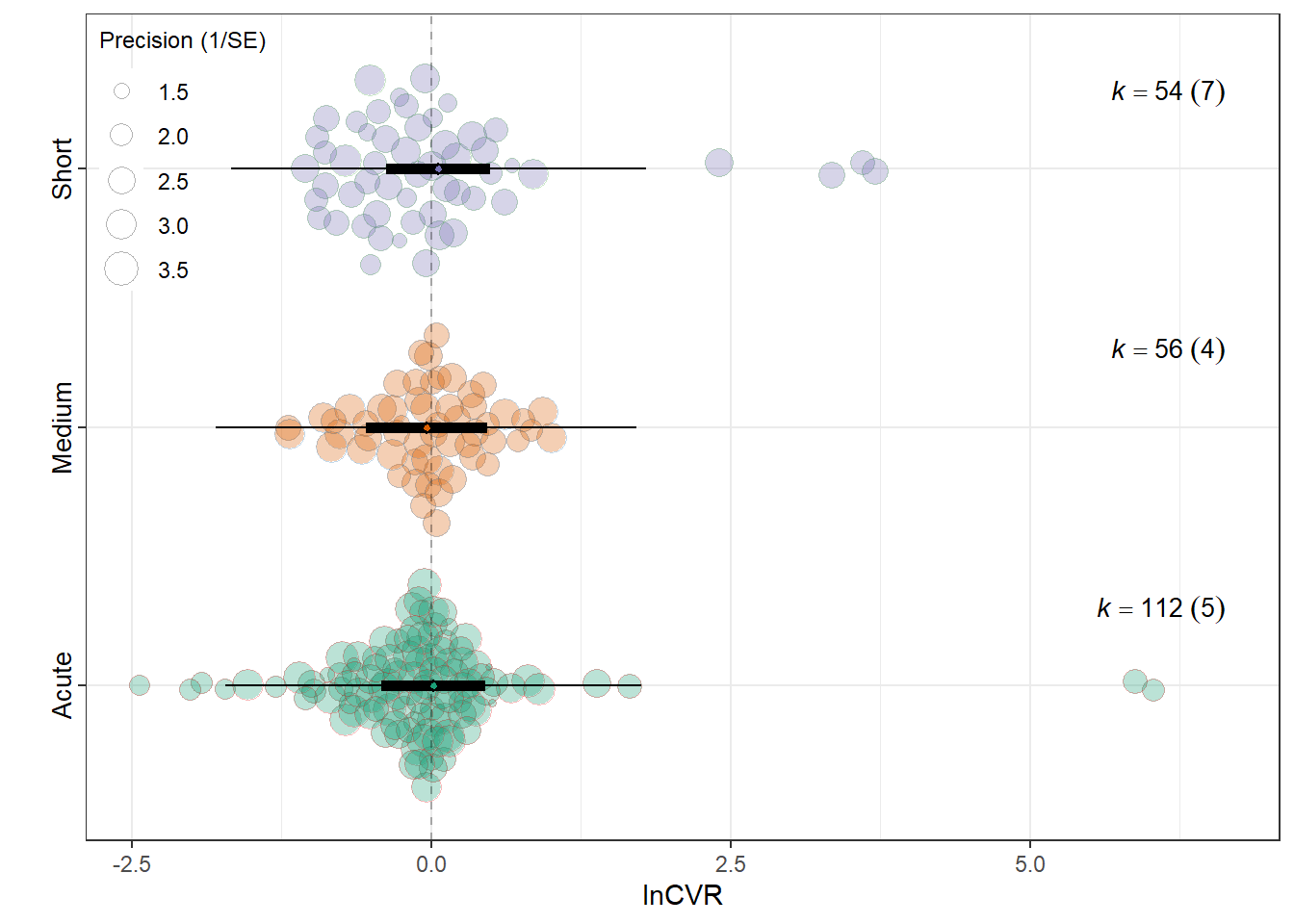
tuk_cvr
Simultaneous Tests for General Linear Hypotheses
Multiple Comparisons of Means: Tukey Contrasts
Fit: metafor::rma.mv(yi = dat[[yi]], V = V, mods = mods_formula, data = dat,
random = list(~1 | Study_ID, ~1 | ES_ID, ~1 | Strain), method = "REML",
test = "t", sparse = TRUE)
Linear Hypotheses:
Estimate Std. Error z value Pr(>|z|)
2 - 1 == 0 -0.06054 0.33837 -0.179 0.858
3 - 1 == 0 0.04097 0.31138 0.132 0.895
3 - 2 == 0 0.10151 0.33811 0.300 0.764
(Adjusted p values reported -- none method)Experimental procedure
MOD <- "Experimental_procedures"
MODS <- "~ Experimental_procedures"
f_rr <- filter_low_k_single(db_rr, VCV_rr, MOD, min_k = 5)
f_vr <- filter_low_k_single(db_vr, VCV_vr, MOD, min_k = 5)
f_cvr <- filter_low_k_single(db_cvr, VCV_cvr, MOD, min_k = 5)
res_rr <- fit_unimod(f_rr$data, f_rr$V, "lnRR", MODS, MOD, expression(lnRR))
res_vr <- fit_unimod(f_vr$data, f_vr$V, "lnVR", MODS, MOD, expression(lnVR))
res_cvr <- fit_unimod(f_cvr$data, f_cvr$V, "lnCVR", MODS, MOD, expression(lnCVR))lnRR
summary(res_rr$model)
Multivariate Meta-Analysis Model (k = 298; method: REML)
logLik Deviance AIC BIC AICc
-241.9324 483.8649 493.8649 512.3167 494.0718
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0546 0.2337 20 no Study_ID
sigma^2.2 0.2291 0.4786 298 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 296) = 5737.2572, p-val < .0001
Test of Moderators (coefficient 2):
F(df1 = 1, df2 = 296) = 0.6644, p-val = 0.4157
Model Results:
estimate se tval df pval ci.lb
intrcpt -0.2227 0.0696 -3.1977 296 0.0015 -0.3597
Experimental_proceduresSham 0.0892 0.1094 0.8151 296 0.4157 -0.1262
ci.ub
intrcpt -0.0856 **
Experimental_proceduresSham 0.3045
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_rr$r2 R2_marginal R2_conditional
0.0045 0.1961 res_rr$plot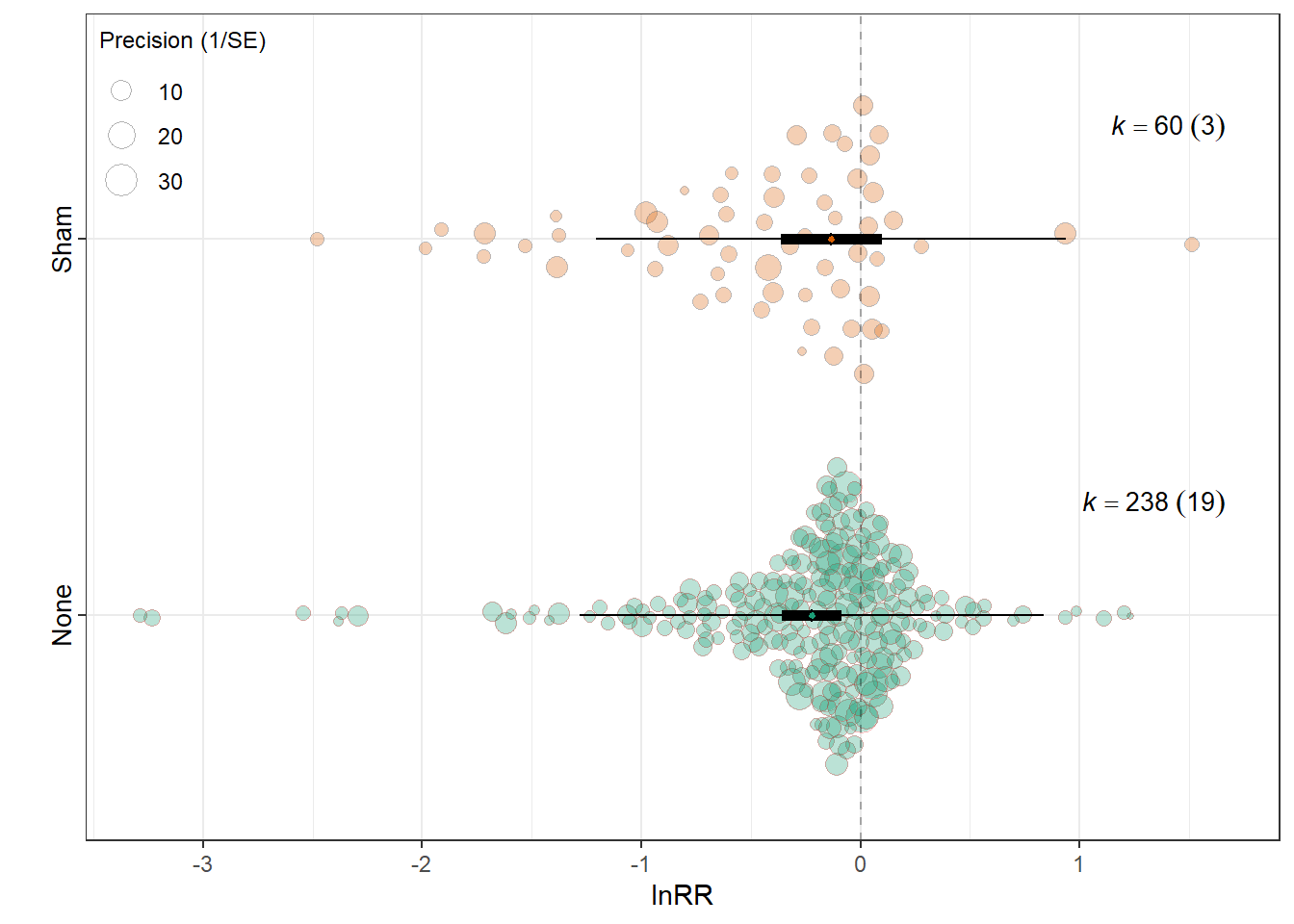
lnVR
summary(res_vr$model)
Multivariate Meta-Analysis Model (k = 222; method: REML)
logLik Deviance AIC BIC AICc
-346.5640 693.1281 703.1281 720.0962 703.4085
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0901 0.3001 16 no Study_ID
sigma^2.2 1.2265 1.1075 222 no ES_ID
sigma^2.3 0.0000 0.0001 6 no Strain
Test for Residual Heterogeneity:
QE(df = 220) = 4386.6575, p-val < .0001
Test of Moderators (coefficient 2):
F(df1 = 1, df2 = 220) = 0.2591, p-val = 0.6112
Model Results:
estimate se tval df pval ci.lb
intrcpt 0.1558 0.1362 1.1438 220 0.2539 -0.1126
Experimental_proceduresSham -0.1239 0.2434 -0.5090 220 0.6112 -0.6036
ci.ub
intrcpt 0.4242
Experimental_proceduresSham 0.3558
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_vr$r2 R2_marginal R2_conditional
0.0021 0.0704 res_vr$plot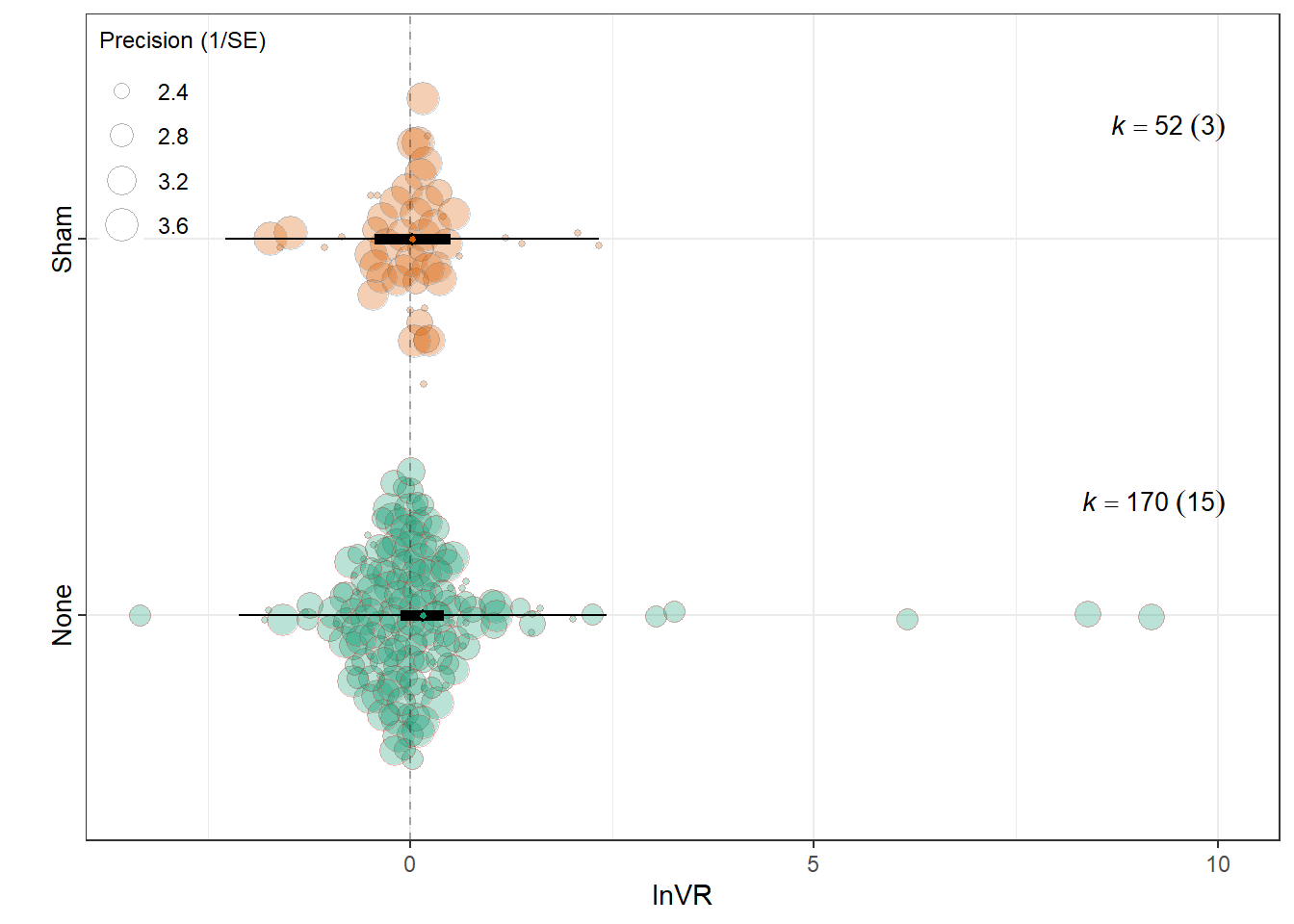
lnCVR
summary(res_cvr$model)
Multivariate Meta-Analysis Model (k = 222; method: REML)
logLik Deviance AIC BIC AICc
-282.4682 564.9365 574.9365 591.9046 575.2168
Variance Components:
estim sqrt nlvls fixed factor
sigma^2.1 0.0911 0.3019 16 no Study_ID
sigma^2.2 0.5936 0.7705 222 no ES_ID
sigma^2.3 0.0000 0.0000 6 no Strain
Test for Residual Heterogeneity:
QE(df = 220) = 1584.4200, p-val < .0001
Test of Moderators (coefficient 2):
F(df1 = 1, df2 = 220) = 0.0651, p-val = 0.7989
Model Results:
estimate se tval df pval ci.lb
intrcpt 0.0246 0.1252 0.1963 220 0.8446 -0.2221
Experimental_proceduresSham -0.0492 0.1927 -0.2551 220 0.7989 -0.4289
ci.ub
intrcpt 0.2712
Experimental_proceduresSham 0.3306
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1res_cvr$r2 R2_marginal R2_conditional
0.0006 0.1336 res_cvr$plot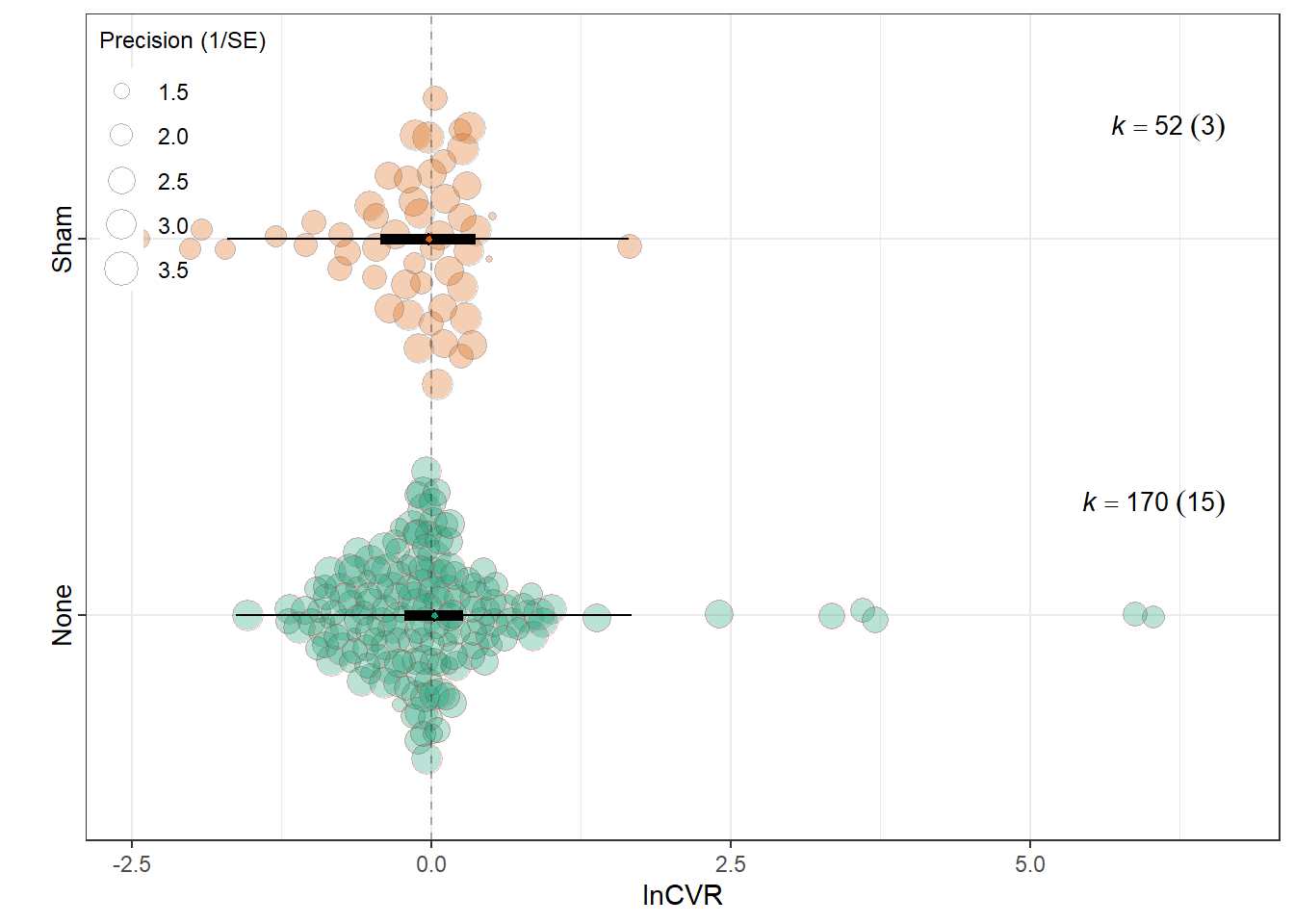
Note
sessionInfo()R version 4.5.2 (2025-10-31 ucrt)
Platform: x86_64-w64-mingw32/x64
Running under: Windows 11 x64 (build 26200)
Matrix products: default
LAPACK version 3.12.1
locale:
[1] LC_COLLATE=English_Guernsey.utf8 LC_CTYPE=English_Guernsey.utf8
[3] LC_MONETARY=English_Guernsey.utf8 LC_NUMERIC=C
[5] LC_TIME=English_Guernsey.utf8
time zone: America/Edmonton
tzcode source: internal
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] multcomp_1.4-29 TH.data_1.1-5 MASS_7.3-65
[4] survival_3.8-6 mvtnorm_1.3-3 orchaRd_2.1.3
[7] metafor_4.8-0 numDeriv_2016.8-1.1 metadat_1.4-0
[10] Matrix_1.7-4 patchwork_1.3.2 lubridate_1.9.4
[13] forcats_1.0.1 stringr_1.6.0 dplyr_1.1.4
[16] purrr_1.2.1 readr_2.1.6 tidyr_1.3.2
[19] tibble_3.3.1 ggplot2_4.0.1 tidyverse_2.0.0
[22] knitr_1.51 here_1.0.2 dtplyr_1.3.2
[25] DT_0.34.0
loaded via a namespace (and not attached):
[1] beeswarm_0.4.0 gtable_0.3.6 xfun_0.56
[4] htmlwidgets_1.6.4 lattice_0.22-7 mathjaxr_2.0-0
[7] tzdb_0.5.0 vctrs_0.7.1 tools_4.5.2
[10] generics_0.1.4 parallel_4.5.2 sandwich_3.1-1
[13] pacman_0.5.1 pkgconfig_2.0.3 data.table_1.18.2.1
[16] RColorBrewer_1.1-3 S7_0.2.1 lifecycle_1.0.5
[19] compiler_4.5.2 farver_2.1.2 codetools_0.2-20
[22] vipor_0.4.7 htmltools_0.5.9 yaml_2.3.12
[25] crayon_1.5.3 pillar_1.11.1 nlme_3.1-168
[28] tidyselect_1.2.1 digest_0.6.39 stringi_1.8.7
[31] labeling_0.4.3 splines_4.5.2 latex2exp_0.9.8
[34] rprojroot_2.1.1 fastmap_1.2.0 grid_4.5.2
[37] cli_3.6.5 magrittr_2.0.4 withr_3.0.2
[40] scales_1.4.0 bit64_4.6.0-1 ggbeeswarm_0.7.3
[43] estimability_1.5.1 timechange_0.3.0 rmarkdown_2.30
[46] emmeans_2.0.1 bit_4.6.0 otel_0.2.0
[49] zoo_1.8-15 hms_1.1.4 coda_0.19-4.1
[52] evaluate_1.0.5 rlang_1.1.7 xtable_1.8-4
[55] glue_1.8.0 rstudioapi_0.18.0 vroom_1.7.0
[58] jsonlite_2.0.0 R6_2.6.1